Probe CUST_6395_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_6395_PI426222305 | JHI_St_60k_v1 | DMT400014382 | TCATTCCAAATTGGGCAACAGGTATATTTCTTCTTTCAAATAAACCTGGCTGCTACCTCT |
All Microarray Probes Designed to Gene DMG400005641
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_6395_PI426222305 | JHI_St_60k_v1 | DMT400014382 | TCATTCCAAATTGGGCAACAGGTATATTTCTTCTTTCAAATAAACCTGGCTGCTACCTCT |
| CUST_6673_PI426222305 | JHI_St_60k_v1 | DMT400014383 | CTCAAGGAGAGAGTAGTACTCCTTTCCGAGACTTTTATTTGTTCAGTTTAGAAAATAAGA |
Microarray Signals from CUST_6395_PI426222305
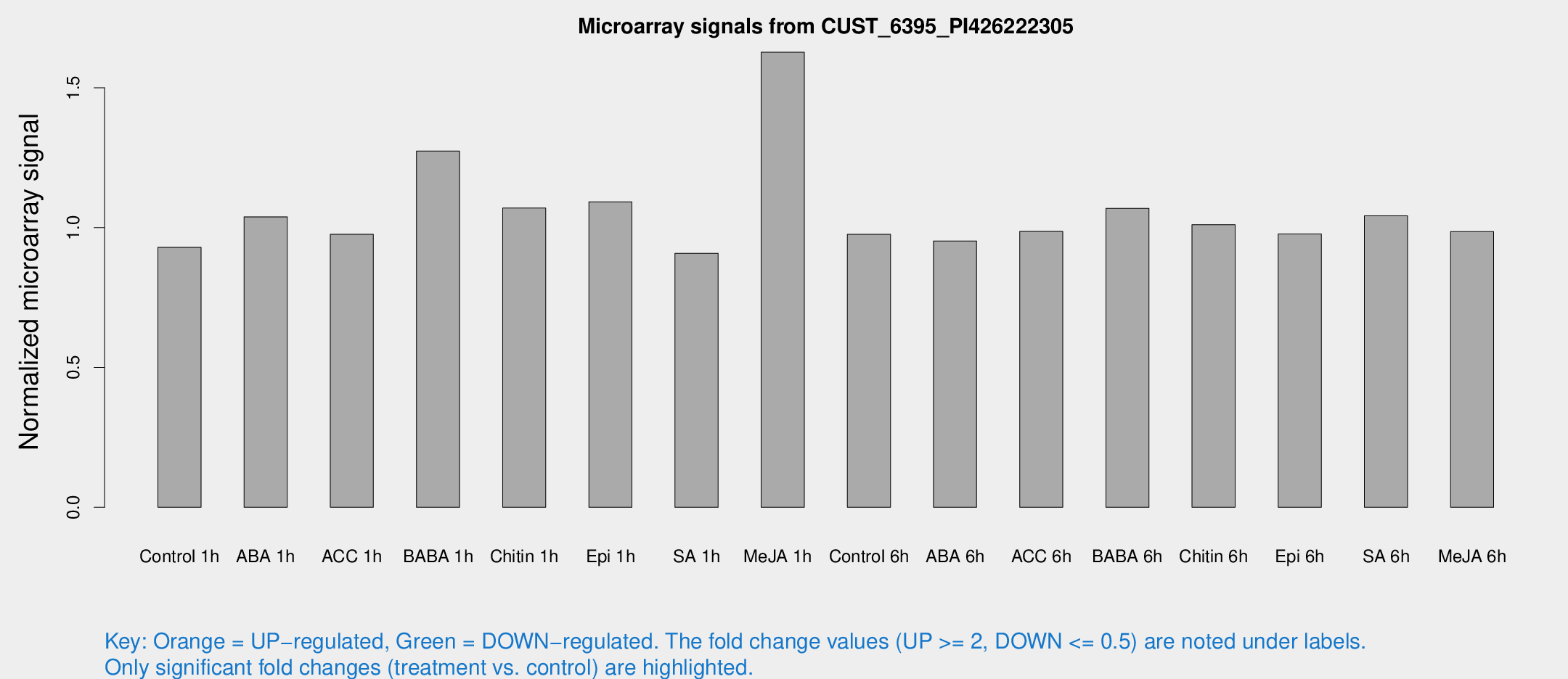
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 5.06223 | 2.93666 | 0.929838 | 0.533555 |
| ABA 1h | 4.93807 | 2.86988 | 1.0388 | 0.589308 |
| ACC 1h | 5.51792 | 3.19806 | 0.97632 | 0.565379 |
| BABA 1h | 7.2374 | 3.15154 | 1.27372 | 0.625338 |
| Chitin 1h | 5.14918 | 2.99451 | 1.07025 | 0.601068 |
| Epi 1h | 5.11333 | 2.96641 | 1.09249 | 0.621216 |
| SA 1h | 5.13161 | 2.97332 | 0.908382 | 0.526289 |
| Me-JA 1h | 8.05099 | 3.08909 | 1.62733 | 0.76253 |
| Control 6h | 5.38025 | 3.11746 | 0.975999 | 0.565378 |
| ABA 6h | 5.5742 | 3.23261 | 0.952548 | 0.551849 |
| ACC 6h | 6.36137 | 3.76925 | 0.986678 | 0.571356 |
| BABA 6h | 6.70419 | 3.53046 | 1.069 | 0.574824 |
| Chitin 6h | 5.91684 | 3.42642 | 1.01071 | 0.585223 |
| Epi 6h | 6.10993 | 3.56704 | 0.977611 | 0.566092 |
| SA 6h | 5.67246 | 3.28692 | 1.04223 | 0.603542 |
| Me-JA 6h | 5.40471 | 3.13224 | 0.98618 | 0.571057 |
Source Transcript PGSC0003DMT400014382 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G68320.1 | +2 | 5e-42 | 142 | 65/79 (82%) | myb domain protein 62 | chr1:25603842-25604884 FORWARD LENGTH=286 |
