Probe CUST_6361_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_6361_PI426222305 | JHI_St_60k_v1 | DMT400036852 | CGATGTTGTGATGAGTAATTTGAGAGGTGAAACAAGAGCAGAGTTAGAGATGATTGAGGA |
All Microarray Probes Designed to Gene DMG400014210
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_6814_PI426222305 | JHI_St_60k_v1 | DMT400036851 | CGTAGTGGGTATATAATCAACTTGTAAAAATGGTCCCTCTTTTGTACCAATTACCTGAAT |
| CUST_6361_PI426222305 | JHI_St_60k_v1 | DMT400036852 | CGATGTTGTGATGAGTAATTTGAGAGGTGAAACAAGAGCAGAGTTAGAGATGATTGAGGA |
Microarray Signals from CUST_6361_PI426222305
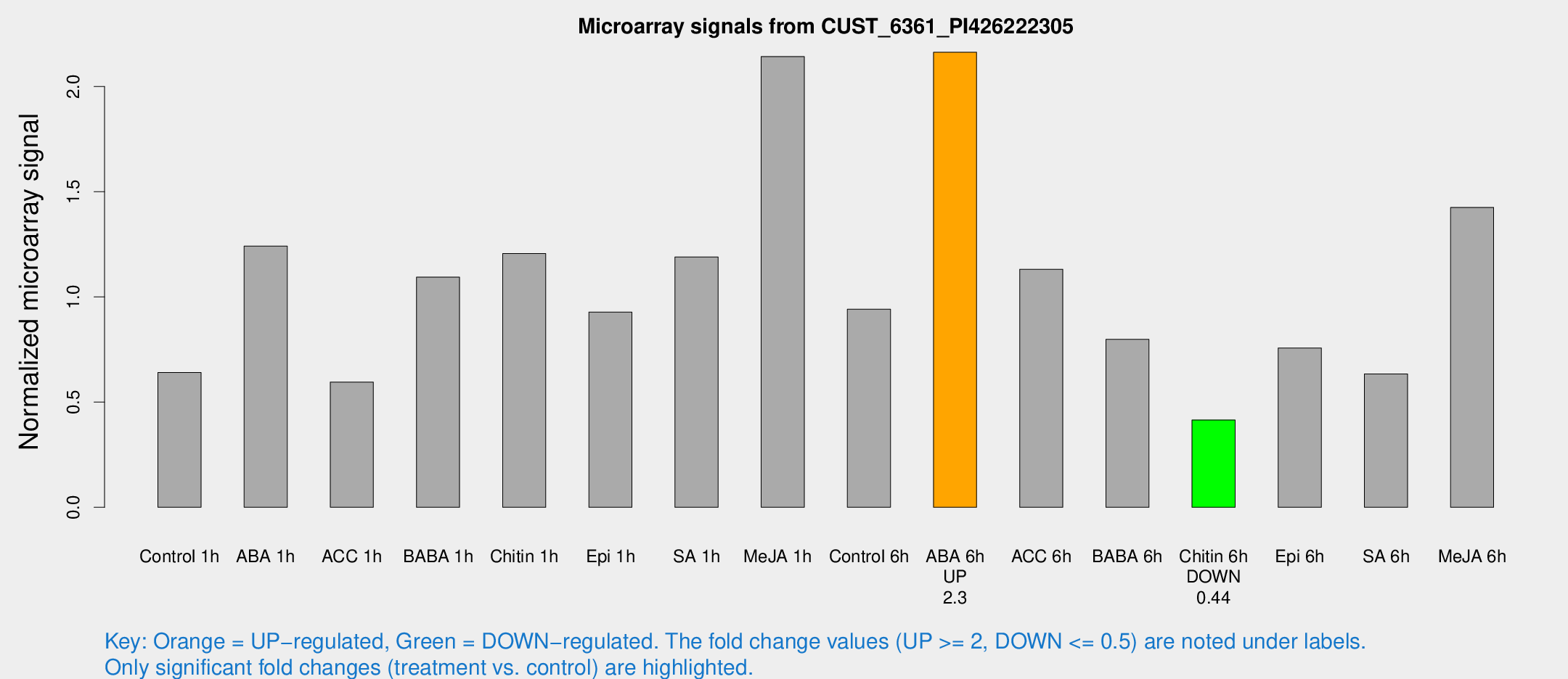
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 2824.88 | 478.634 | 0.640584 | 0.140418 |
| ABA 1h | 5174.13 | 1673.29 | 1.2417 | 0.30294 |
| ACC 1h | 2690.81 | 436.643 | 0.595393 | 0.116347 |
| BABA 1h | 5119.4 | 1670.98 | 1.09403 | 0.313384 |
| Chitin 1h | 4656.16 | 269.363 | 1.20609 | 0.139571 |
| Epi 1h | 3769.31 | 1017.37 | 0.927857 | 0.29578 |
| SA 1h | 5318.54 | 673.888 | 1.1901 | 0.0818303 |
| Me-JA 1h | 7669.7 | 1148.2 | 2.14272 | 0.140284 |
| Control 6h | 4122.18 | 538.575 | 0.941687 | 0.0543738 |
| ABA 6h | 10378.2 | 2357.5 | 2.16307 | 0.380479 |
| ACC 6h | 5576.36 | 322.85 | 1.13137 | 0.0981324 |
| BABA 6h | 3889.23 | 494.76 | 0.798391 | 0.0850161 |
| Chitin 6h | 1894.73 | 109.568 | 0.414927 | 0.0265957 |
| Epi 6h | 3818.64 | 815.797 | 0.75677 | 0.145292 |
| SA 6h | 3281.84 | 1162.93 | 0.634192 | 0.26651 |
| Me-JA 6h | 6507.93 | 1686.24 | 1.42526 | 0.332858 |
Source Transcript PGSC0003DMT400036852 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G80920.1 | +3 | 2e-53 | 174 | 87/142 (61%) | Chaperone DnaJ-domain superfamily protein | chr1:30403863-30404549 REVERSE LENGTH=163 |
