Probe CUST_5892_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_5892_PI426222305 | JHI_St_60k_v1 | DMT400022943 | TTGGGGTGGATGTGTCTCCATTTTTTGGATCTTGATGATGTAAAAGTGTATAATTTCATC |
All Microarray Probes Designed to Gene DMG401008889
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_5524_PI426222305 | JHI_St_60k_v1 | DMT400022947 | ATTCTACATCAGGTGCATCCTTTAAGGCTATTGGGCTCGATCGAACCAGGAATTACGTGT |
| CUST_5892_PI426222305 | JHI_St_60k_v1 | DMT400022943 | TTGGGGTGGATGTGTCTCCATTTTTTGGATCTTGATGATGTAAAAGTGTATAATTTCATC |
Microarray Signals from CUST_5892_PI426222305
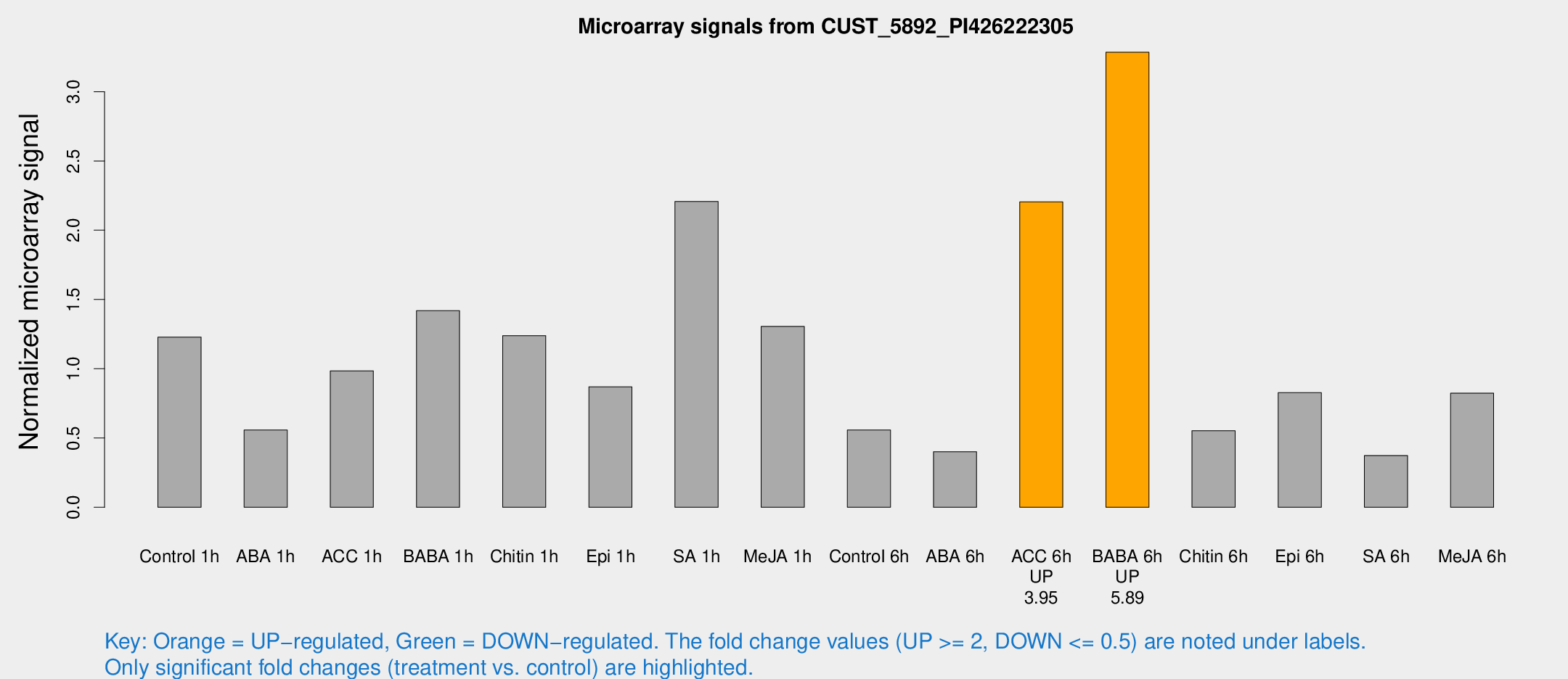
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 220.398 | 64.2105 | 1.22889 | 0.269031 |
| ABA 1h | 85.0986 | 15.9685 | 0.55766 | 0.101788 |
| ACC 1h | 194.358 | 60.8698 | 0.985161 | 0.361832 |
| BABA 1h | 237.981 | 48.2853 | 1.41903 | 0.176187 |
| Chitin 1h | 185.405 | 11.3815 | 1.23937 | 0.0801048 |
| Epi 1h | 125.74 | 9.23943 | 0.869858 | 0.0564979 |
| SA 1h | 383.29 | 49.3419 | 2.20761 | 0.274729 |
| Me-JA 1h | 181.527 | 29.2576 | 1.30591 | 0.144747 |
| Control 6h | 95.5221 | 15.5501 | 0.558027 | 0.0577227 |
| ABA 6h | 77.1527 | 21.1449 | 0.4014 | 0.10293 |
| ACC 6h | 435.382 | 77.8081 | 2.20439 | 0.350177 |
| BABA 6h | 622.445 | 87.8524 | 3.2848 | 0.380136 |
| Chitin 6h | 100.474 | 15.6788 | 0.55297 | 0.0919522 |
| Epi 6h | 159.878 | 27.9133 | 0.827572 | 0.121864 |
| SA 6h | 66.6096 | 16.9023 | 0.373907 | 0.146372 |
| Me-JA 6h | 151.064 | 48.0686 | 0.824099 | 0.204078 |
Source Transcript PGSC0003DMT400022943 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G37820.1 | +1 | 5e-11 | 61 | 46/163 (28%) | Cysteine/Histidine-rich C1 domain family protein | chr2:15847541-15848413 FORWARD LENGTH=290 |
