Probe CUST_5838_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_5838_PI426222305 | JHI_St_60k_v1 | DMT400022761 | CTGTCACAGCTTAGAATCAATAGTAGTACTATATCATCATCATGATGTTGGCAGAAGAAA |
All Microarray Probes Designed to Gene DMG400008829
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_5838_PI426222305 | JHI_St_60k_v1 | DMT400022761 | CTGTCACAGCTTAGAATCAATAGTAGTACTATATCATCATCATGATGTTGGCAGAAGAAA |
Microarray Signals from CUST_5838_PI426222305
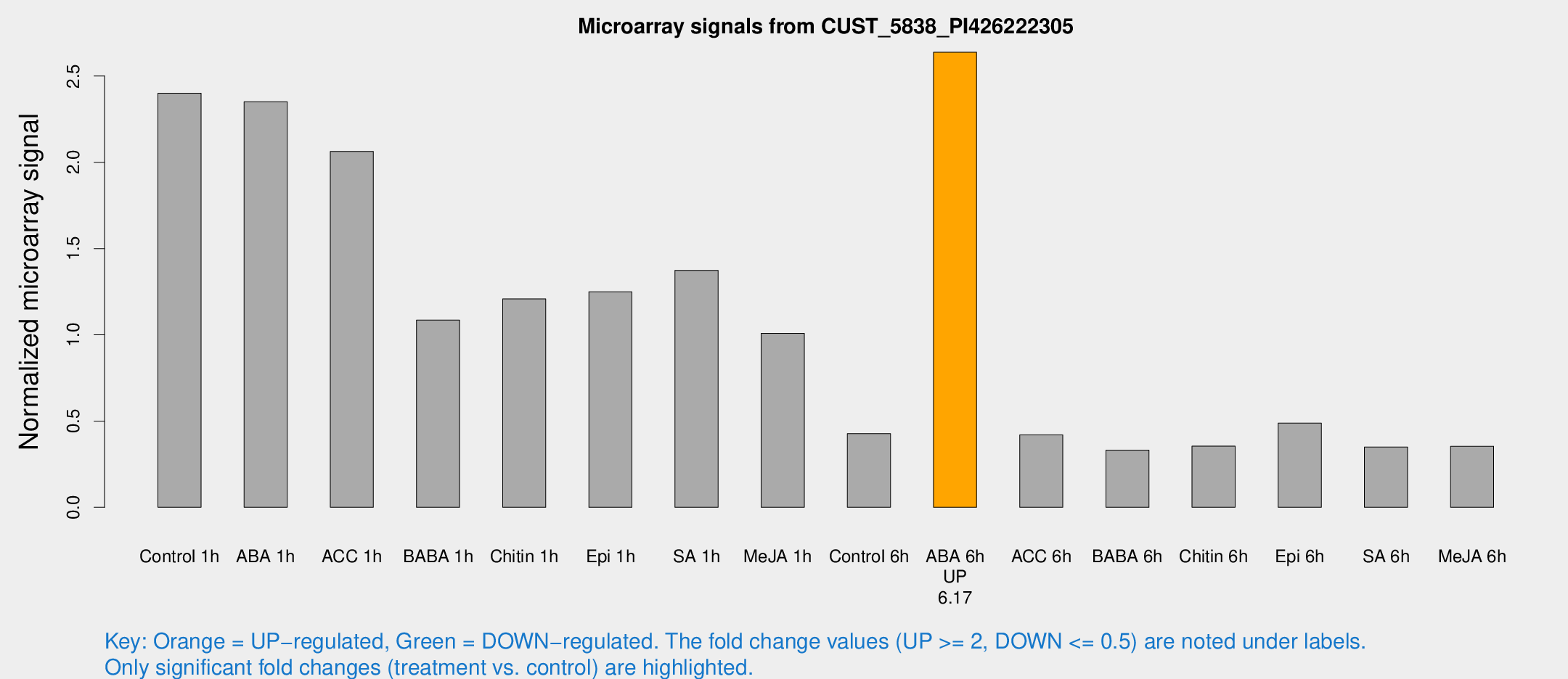
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 886.73 | 156.479 | 2.40075 | 0.25556 |
| ABA 1h | 751.223 | 64.2797 | 2.35155 | 0.371464 |
| ACC 1h | 762.318 | 46.3216 | 2.06312 | 0.146622 |
| BABA 1h | 382.862 | 54.2775 | 1.0849 | 0.0685246 |
| Chitin 1h | 397.634 | 56.5422 | 1.20787 | 0.146665 |
| Epi 1h | 394.425 | 49.632 | 1.24916 | 0.147375 |
| SA 1h | 506.28 | 29.4323 | 1.37344 | 0.0797748 |
| Me-JA 1h | 304.685 | 56.3977 | 1.00851 | 0.190961 |
| Control 6h | 171.701 | 49.8416 | 0.427251 | 0.117332 |
| ABA 6h | 1008.23 | 58.488 | 2.63816 | 0.152628 |
| ACC 6h | 174.982 | 19.9545 | 0.420118 | 0.0278568 |
| BABA 6h | 137.098 | 22.081 | 0.331614 | 0.0632828 |
| Chitin 6h | 138.522 | 21.2873 | 0.354946 | 0.0576154 |
| Epi 6h | 200.112 | 23.7079 | 0.487953 | 0.0852183 |
| SA 6h | 124.632 | 9.74034 | 0.349175 | 0.0624583 |
| Me-JA 6h | 126.524 | 8.1511 | 0.353555 | 0.0330308 |
Source Transcript PGSC0003DMT400022761 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G02230.1 | +1 | 2e-135 | 394 | 199/294 (68%) | Haloacid dehalogenase-like hydrolase (HAD) superfamily protein | chr5:449132-450507 FORWARD LENGTH=280 |
