Probe CUST_5784_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_5784_PI426222305 | JHI_St_60k_v1 | DMT400022974 | CTCATCGAAGCAGCGAAGAAAAATCCACTAACTAAAGAACGTATGGAAAAAAGTATTAGT |
All Microarray Probes Designed to Gene DMG402008895
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_5784_PI426222305 | JHI_St_60k_v1 | DMT400022974 | CTCATCGAAGCAGCGAAGAAAAATCCACTAACTAAAGAACGTATGGAAAAAAGTATTAGT |
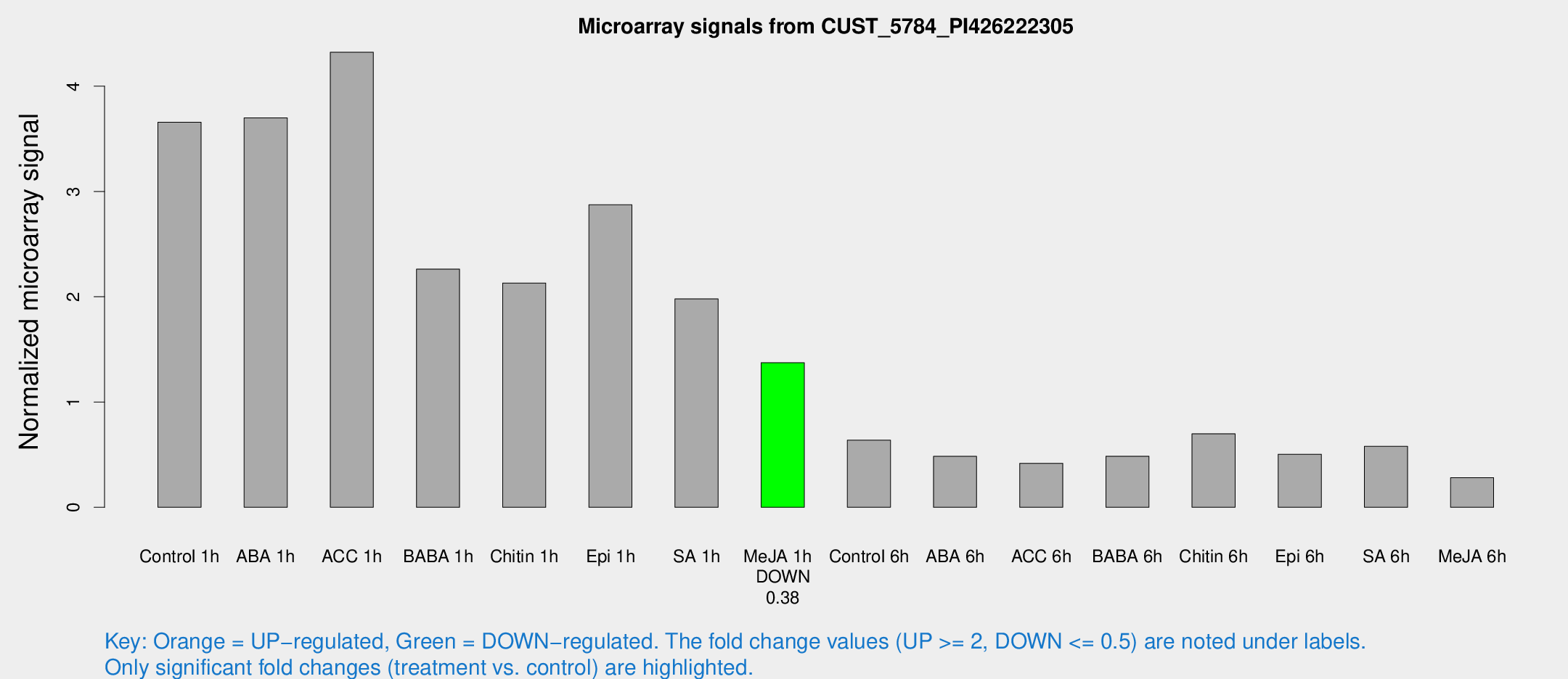
Microarray Signals from CUST_5784_PI426222305
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 485.669 | 93.9306 | 3.65772 | 0.446433 |
| ABA 1h | 421.85 | 31.7322 | 3.69969 | 0.513009 |
| ACC 1h | 575.198 | 56.9521 | 4.32285 | 0.251019 |
| BABA 1h | 295.28 | 62.9787 | 2.2628 | 0.319628 |
| Chitin 1h | 249.409 | 30.2952 | 2.13025 | 0.209477 |
| Epi 1h | 320.172 | 20.1727 | 2.87362 | 0.17015 |
| SA 1h | 272.913 | 61.1021 | 1.97974 | 0.315511 |
| Me-JA 1h | 143.729 | 8.87585 | 1.37294 | 0.1289 |
| Control 6h | 90.4775 | 24.4899 | 0.637147 | 0.16314 |
| ABA 6h | 67.335 | 9.83417 | 0.484309 | 0.0938036 |
| ACC 6h | 66.7978 | 20.1291 | 0.417825 | 0.0611592 |
| BABA 6h | 71.8609 | 12.57 | 0.485381 | 0.0968692 |
| Chitin 6h | 96.5506 | 10.4856 | 0.698597 | 0.066538 |
| Epi 6h | 83.0673 | 24.9708 | 0.504398 | 0.269717 |
| SA 6h | 74.4277 | 8.85288 | 0.579152 | 0.062713 |
| Me-JA 6h | 39.1431 | 11.3001 | 0.281707 | 0.0752992 |
Source Transcript PGSC0003DMT400022974 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G07440.1 | +2 | 3e-101 | 301 | 153/255 (60%) | NAD(P)-binding Rossmann-fold superfamily protein | chr1:2286436-2287665 REVERSE LENGTH=266 |
