Probe CUST_5622_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_5622_PI426222305 | JHI_St_60k_v1 | DMT400022959 | ACTTTTGCAGTCATCTTTTATTGTGGAGTAGTTAAATACTATGCTCCTGGTGAGGCTATG |
All Microarray Probes Designed to Gene DMG401008890
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_5622_PI426222305 | JHI_St_60k_v1 | DMT400022959 | ACTTTTGCAGTCATCTTTTATTGTGGAGTAGTTAAATACTATGCTCCTGGTGAGGCTATG |
| CUST_5734_PI426222305 | JHI_St_60k_v1 | DMT400022949 | ATTTCATTTGTTCACGAGACGCATGGCCAGTACAGAACACTTGAAGAGCTGTGGGATGAT |
Microarray Signals from CUST_5622_PI426222305
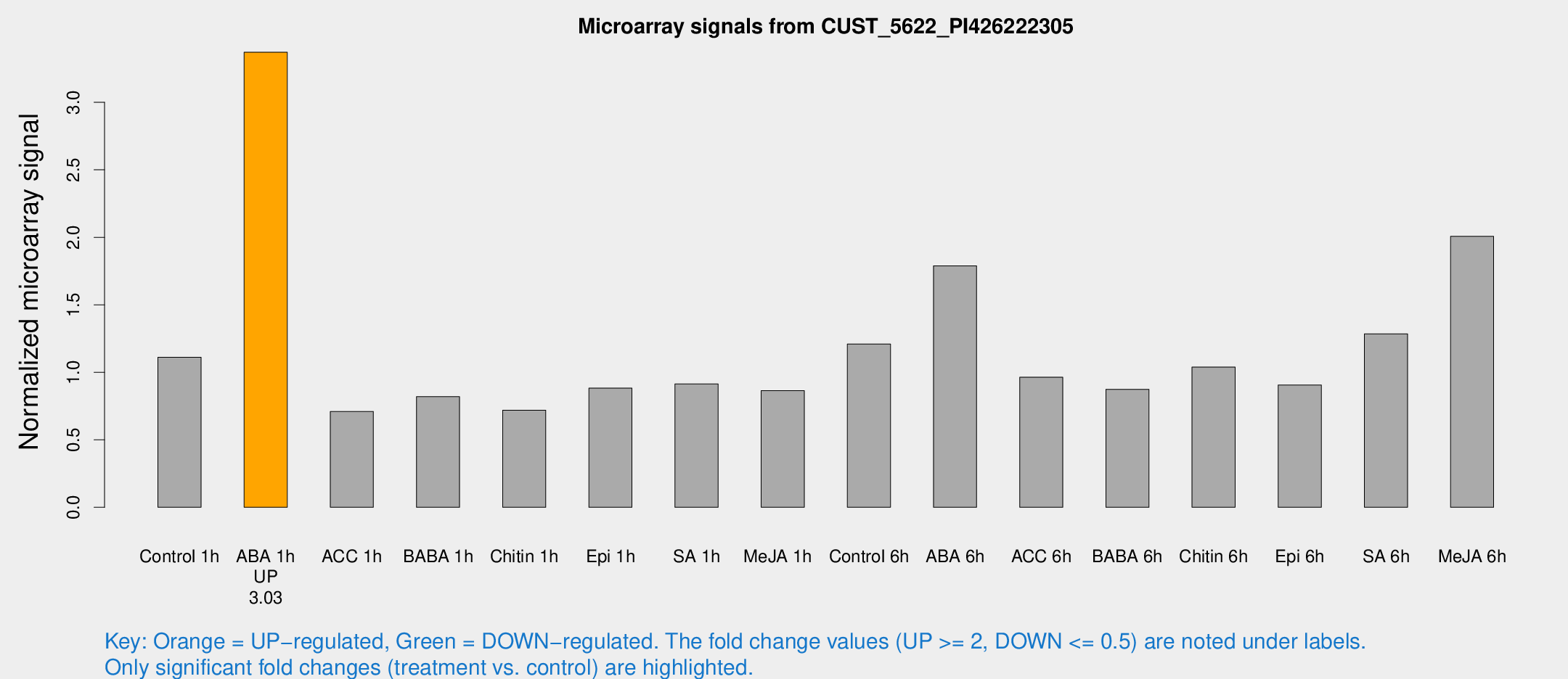
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 252.005 | 18.3423 | 1.11094 | 0.066914 |
| ABA 1h | 676.043 | 45.442 | 3.37126 | 0.232617 |
| ACC 1h | 203.921 | 76.2673 | 0.709691 | 0.360455 |
| BABA 1h | 186.831 | 37.7449 | 0.819552 | 0.101716 |
| Chitin 1h | 147.935 | 16.6819 | 0.719465 | 0.0456359 |
| Epi 1h | 176.662 | 27.1634 | 0.882722 | 0.137917 |
| SA 1h | 223.152 | 52.4842 | 0.913885 | 0.20218 |
| Me-JA 1h | 159.7 | 10.0003 | 0.864591 | 0.0540607 |
| Control 6h | 295.898 | 72.5298 | 1.21003 | 0.25326 |
| ABA 6h | 433.516 | 41.7782 | 1.78899 | 0.122835 |
| ACC 6h | 251.186 | 19.9198 | 0.963875 | 0.11536 |
| BABA 6h | 226.414 | 34.6219 | 0.874233 | 0.163907 |
| Chitin 6h | 253.049 | 30.812 | 1.03886 | 0.0921957 |
| Epi 6h | 232.094 | 19.3647 | 0.905751 | 0.0621647 |
| SA 6h | 287.86 | 17.1677 | 1.2852 | 0.162458 |
| Me-JA 6h | 463.868 | 72.5353 | 2.00708 | 0.407179 |
Source Transcript PGSC0003DMT400022959 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G37790.1 | +1 | 2e-167 | 477 | 228/314 (73%) | NAD(P)-linked oxidoreductase superfamily protein | chr2:15838838-15840752 FORWARD LENGTH=314 |
