Probe CUST_560_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_560_PI426222305 | JHI_St_60k_v1 | DMT400074460 | GCAAATCCTGTAAATTATGTGAGGGTTCAGCTTGGCATCAAAGAGAAGAAGTGGAGGAAG |
All Microarray Probes Designed to Gene DMG400028939
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_560_PI426222305 | JHI_St_60k_v1 | DMT400074460 | GCAAATCCTGTAAATTATGTGAGGGTTCAGCTTGGCATCAAAGAGAAGAAGTGGAGGAAG |
Microarray Signals from CUST_560_PI426222305
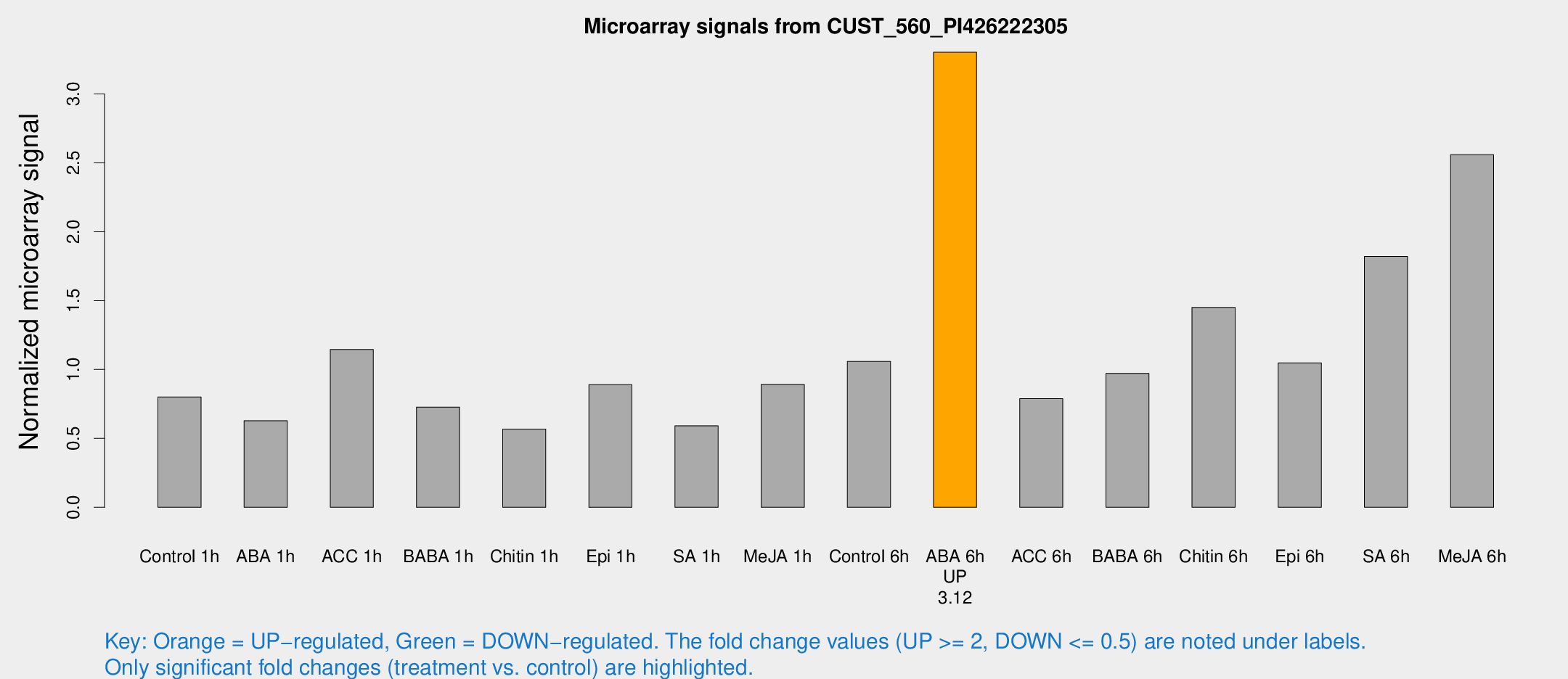
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 10.874 | 3.7601 | 0.800143 | 0.315768 |
| ABA 1h | 7.20621 | 3.65095 | 0.62867 | 0.322481 |
| ACC 1h | 17.4353 | 5.95012 | 1.14492 | 0.636031 |
| BABA 1h | 9.45098 | 4.03874 | 0.727529 | 0.341043 |
| Chitin 1h | 6.58268 | 3.82552 | 0.568042 | 0.329285 |
| Epi 1h | 10.7593 | 3.74928 | 0.889805 | 0.365244 |
| SA 1h | 7.96844 | 3.6953 | 0.591553 | 0.28395 |
| Me-JA 1h | 10.0432 | 3.75843 | 0.891328 | 0.376028 |
| Control 6h | 15.2589 | 4.63251 | 1.05854 | 0.376054 |
| ABA 6h | 48.5638 | 11.7952 | 3.30248 | 0.811111 |
| ACC 6h | 12.879 | 4.86643 | 0.788447 | 0.366902 |
| BABA 6h | 13.991 | 4.38149 | 0.971834 | 0.305566 |
| Chitin 6h | 19.9592 | 4.39201 | 1.45042 | 0.323366 |
| Epi 6h | 17.7737 | 6.07951 | 1.04752 | 0.429837 |
| SA 6h | 23.3676 | 4.29554 | 1.82077 | 0.343905 |
| Me-JA 6h | 32.8366 | 4.23797 | 2.55867 | 0.331042 |
Source Transcript PGSC0003DMT400074460 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | Solyc09g074110.2 | +1 | 2e-22 | 92 | 42/49 (86%) | genomic_reference:SL2.50ch09 gene_region:61291740-61320947 transcript_region:SL2.50ch09:61291740..61320947+ functional_description:AGAP009276-PA (Fragment) (AHRD V1 *--- Q7PVH5_ANOGA) |
| TAIR PP10 | None | - | - | - | - | - |
