Probe CUST_5437_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_5437_PI426222305 | JHI_St_60k_v1 | DMT400009295 | CACACACGGTAGTAATCTTATTAAACAACAGAAGACCTCTTTAGAAACTGATTAGCAACA |
All Microarray Probes Designed to Gene DMG400003607
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_5437_PI426222305 | JHI_St_60k_v1 | DMT400009295 | CACACACGGTAGTAATCTTATTAAACAACAGAAGACCTCTTTAGAAACTGATTAGCAACA |
| CUST_5093_PI426222305 | JHI_St_60k_v1 | DMT400009296 | CAAATTTAATTAGCAAGGGAGAAGTGGAAAAGGAAAAGCTTGATTCTTACGAAGTACACT |
Microarray Signals from CUST_5437_PI426222305
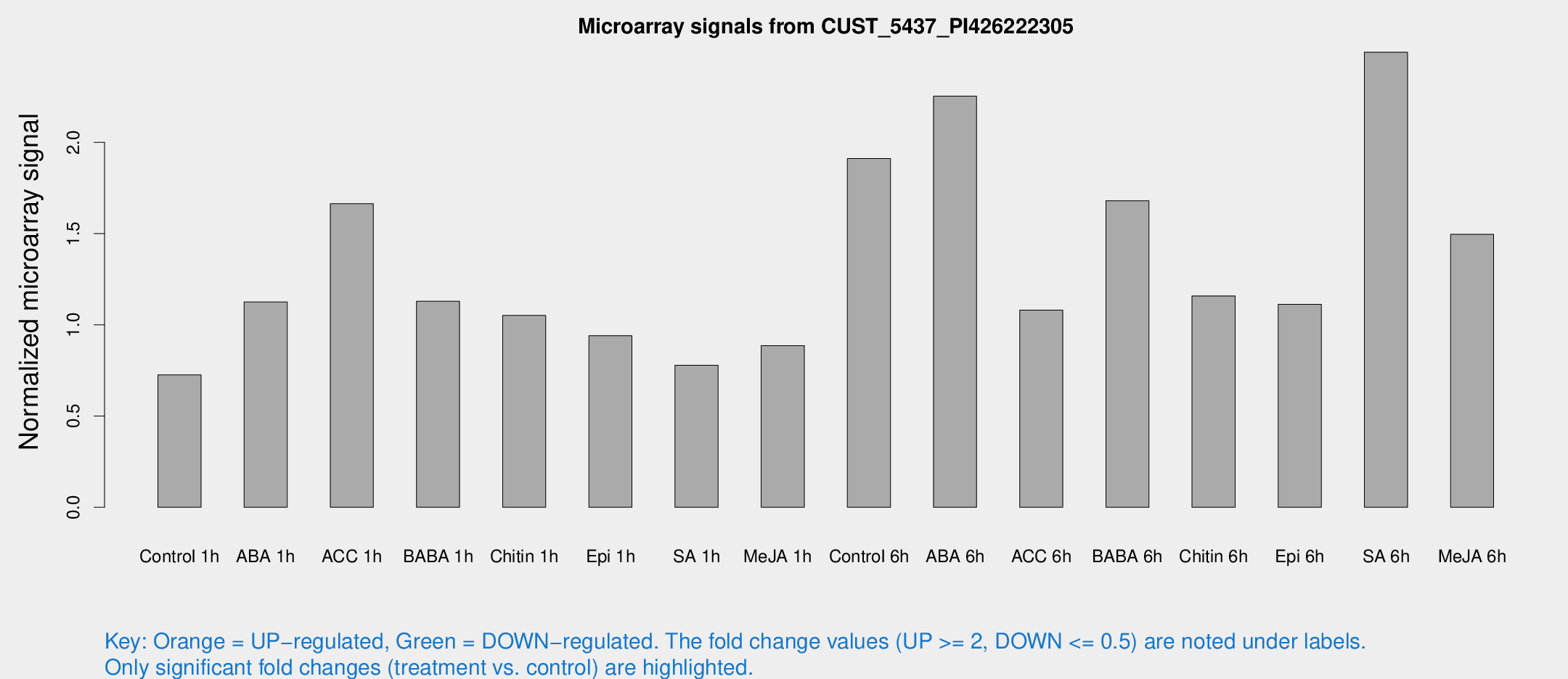
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 6.33023 | 3.58516 | 0.725952 | 0.411063 |
| ABA 1h | 10.2508 | 4.46304 | 1.12521 | 0.524373 |
| ACC 1h | 17.5577 | 6.0623 | 1.66415 | 1.04703 |
| BABA 1h | 9.80368 | 3.86537 | 1.13007 | 0.489958 |
| Chitin 1h | 8.33355 | 3.59643 | 1.05201 | 0.465226 |
| Epi 1h | 7.157 | 3.45488 | 0.940796 | 0.472871 |
| SA 1h | 7.00683 | 3.60008 | 0.778255 | 0.408437 |
| Me-JA 1h | 6.32061 | 3.69152 | 0.886016 | 0.513111 |
| Control 6h | 17.0436 | 3.88198 | 1.91208 | 0.472058 |
| ABA 6h | 26.5304 | 10.2305 | 2.25375 | 1.39621 |
| ACC 6h | 11.2262 | 4.36913 | 1.08128 | 0.456086 |
| BABA 6h | 18.3149 | 5.66894 | 1.68049 | 0.575203 |
| Chitin 6h | 11.5607 | 4.1786 | 1.15844 | 0.500691 |
| Epi 6h | 12.309 | 4.4452 | 1.11276 | 0.50617 |
| SA 6h | 22.0881 | 4.11089 | 2.49442 | 0.506476 |
| Me-JA 6h | 14.3735 | 4.92782 | 1.49624 | 0.492321 |
Source Transcript PGSC0003DMT400009295 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G36470.1 | +1 | 1e-60 | 211 | 100/168 (60%) | S-adenosyl-L-methionine-dependent methyltransferases superfamily protein | chr4:17215128-17216475 REVERSE LENGTH=371 |
