Probe CUST_5310_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_5310_PI426222305 | JHI_St_60k_v1 | DMT400009298 | TCTTCATCGGCCTGTTCCAATTTATGTAACACACTTTCTATGTTATTCTGTCCAAAAAAG |
All Microarray Probes Designed to Gene DMG400003609
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_5310_PI426222305 | JHI_St_60k_v1 | DMT400009298 | TCTTCATCGGCCTGTTCCAATTTATGTAACACACTTTCTATGTTATTCTGTCCAAAAAAG |
Microarray Signals from CUST_5310_PI426222305
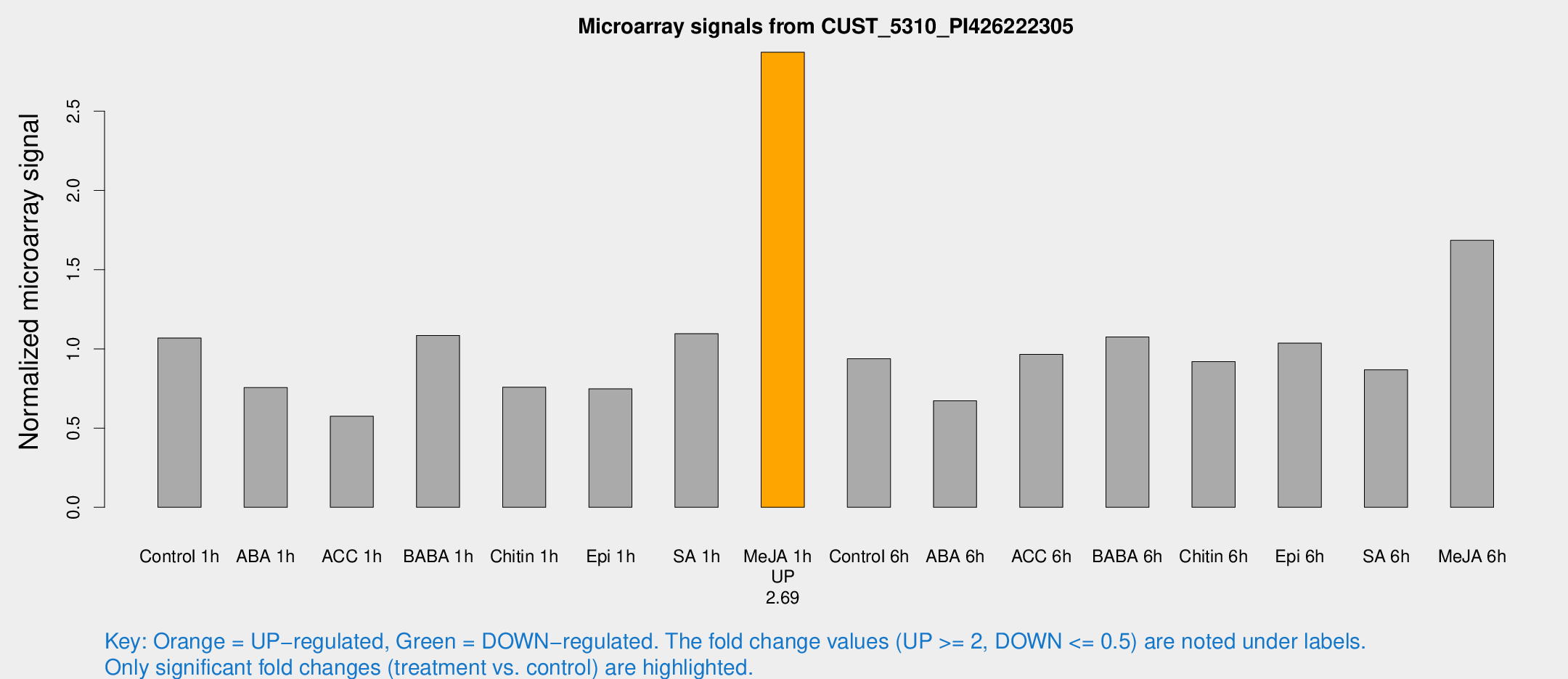
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 95.5578 | 18.6187 | 1.06877 | 0.140339 |
| ABA 1h | 57.9767 | 4.48524 | 0.755429 | 0.0589056 |
| ACC 1h | 75.3969 | 32.4968 | 0.574848 | 0.548997 |
| BABA 1h | 92.2617 | 12.751 | 1.0849 | 0.24873 |
| Chitin 1h | 64.5335 | 18.7842 | 0.75786 | 0.220326 |
| Epi 1h | 58.4471 | 12.4618 | 0.747794 | 0.145654 |
| SA 1h | 104.002 | 24.057 | 1.09579 | 0.388666 |
| Me-JA 1h | 205.377 | 24.6227 | 2.87261 | 0.341277 |
| Control 6h | 90.1937 | 27.9644 | 0.937948 | 0.237195 |
| ABA 6h | 65.0122 | 15.5625 | 0.672107 | 0.190399 |
| ACC 6h | 99.1882 | 19.7545 | 0.9652 | 0.0699142 |
| BABA 6h | 104.67 | 8.76619 | 1.07511 | 0.0929167 |
| Chitin 6h | 88.7734 | 19.2619 | 0.919384 | 0.199713 |
| Epi 6h | 105.977 | 22.1607 | 1.03629 | 0.335593 |
| SA 6h | 74.6571 | 6.16665 | 0.86771 | 0.159467 |
| Me-JA 6h | 149.045 | 23.526 | 1.68525 | 0.421102 |
Source Transcript PGSC0003DMT400009298 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G27660.1 | +2 | 3e-22 | 100 | 53/97 (55%) | basic helix-loop-helix (bHLH) DNA-binding superfamily protein | chr1:9621701-9625666 FORWARD LENGTH=453 |
