Probe CUST_5277_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_5277_PI426222305 | JHI_St_60k_v1 | DMT400009232 | AGTTTGGCAACTCATCTATTTTCTACTATCCTCGTTTTTCTCATACAGGTTCCCGATGAA |
All Microarray Probes Designed to Gene DMG400003585
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_5090_PI426222305 | JHI_St_60k_v1 | DMT400009233 | GATTGGATGAAGTTGATGAGGCCACTAAATGAAGAGAAAGATCACTGGGTTCCCGATGAA |
| CUST_5277_PI426222305 | JHI_St_60k_v1 | DMT400009232 | AGTTTGGCAACTCATCTATTTTCTACTATCCTCGTTTTTCTCATACAGGTTCCCGATGAA |
Microarray Signals from CUST_5277_PI426222305
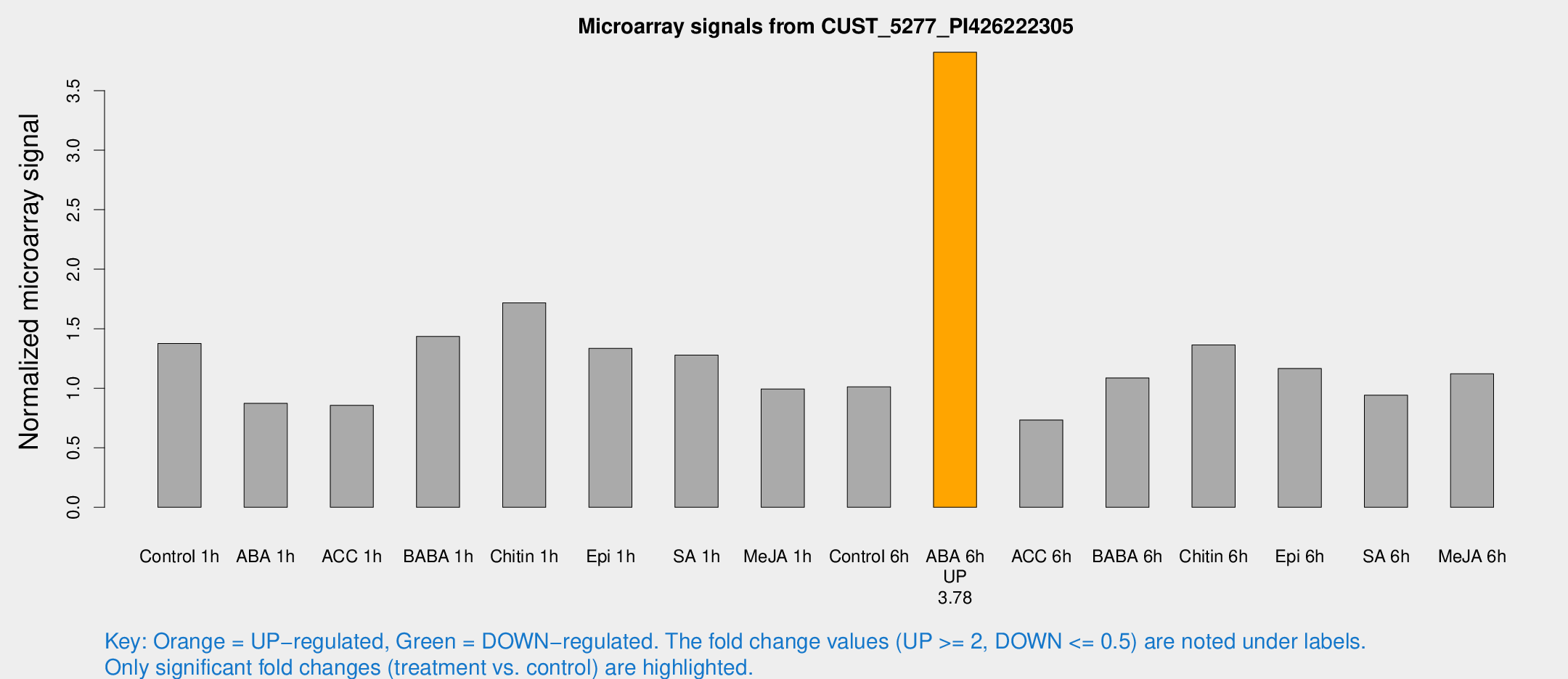
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 11.4169 | 3.20984 | 1.37642 | 0.42908 |
| ABA 1h | 6.32697 | 3.06397 | 0.872816 | 0.444414 |
| ACC 1h | 7.22909 | 3.45559 | 0.856894 | 0.43328 |
| BABA 1h | 14.0736 | 6.99945 | 1.43488 | 0.676643 |
| Chitin 1h | 12.4072 | 3.27976 | 1.71767 | 0.4574 |
| Epi 1h | 9.85946 | 3.16592 | 1.33497 | 0.511119 |
| SA 1h | 11.7419 | 4.10173 | 1.27788 | 0.548217 |
| Me-JA 1h | 6.85389 | 3.26468 | 0.994435 | 0.507146 |
| Control 6h | 8.95681 | 3.41225 | 1.01141 | 0.476232 |
| ABA 6h | 33.6956 | 5.92108 | 3.82249 | 0.563087 |
| ACC 6h | 6.8642 | 4.01105 | 0.733608 | 0.418711 |
| BABA 6h | 11.1038 | 4.17331 | 1.08659 | 0.46387 |
| Chitin 6h | 14.1251 | 6.35619 | 1.36344 | 0.642066 |
| Epi 6h | 11.8249 | 4.24793 | 1.16636 | 0.486445 |
| SA 6h | 7.55003 | 3.51324 | 0.942365 | 0.450916 |
| Me-JA 6h | 9.2797 | 3.39766 | 1.12109 | 0.455324 |
Source Transcript PGSC0003DMT400009232 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G20110.1 | +3 | 2e-82 | 278 | 195/456 (43%) | RING/FYVE/PHD zinc finger superfamily protein | chr1:6971554-6974578 FORWARD LENGTH=601 |
