Probe CUST_52580_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_52580_PI426222305 | JHI_St_60k_v1 | DMT400017286 | CATGATGGAATTAGAGAAGCAAGGGAAAATAGTACCATGGTGTTCACAACTTGAAGTCTT |
All Microarray Probes Designed to Gene DMG400006737
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_52580_PI426222305 | JHI_St_60k_v1 | DMT400017286 | CATGATGGAATTAGAGAAGCAAGGGAAAATAGTACCATGGTGTTCACAACTTGAAGTCTT |
Microarray Signals from CUST_52580_PI426222305
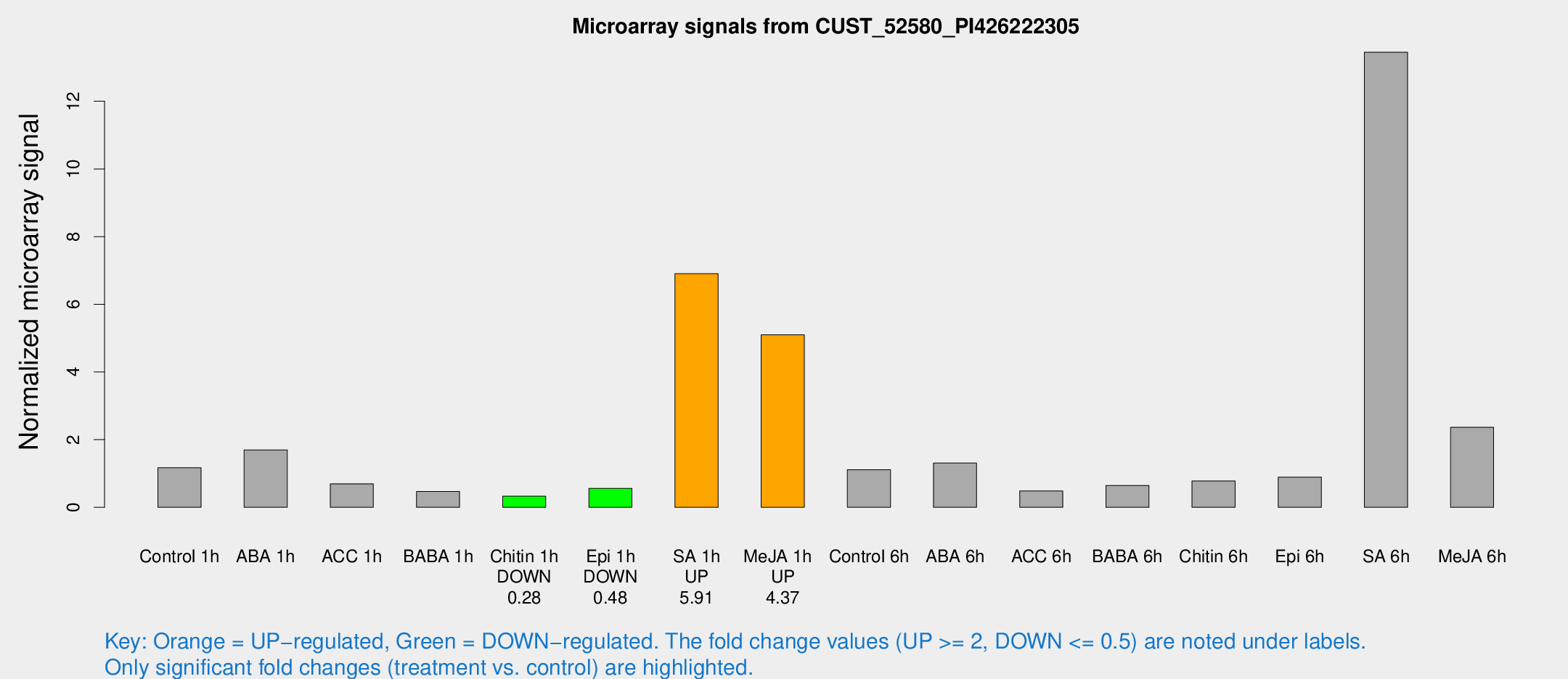
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 78.7886 | 16.0991 | 1.16753 | 0.172521 |
| ABA 1h | 100.605 | 16.9378 | 1.69494 | 0.240593 |
| ACC 1h | 48.7026 | 11.2556 | 0.693243 | 0.126375 |
| BABA 1h | 37.1983 | 14.7152 | 0.466039 | 0.241315 |
| Chitin 1h | 19.2702 | 3.2558 | 0.327266 | 0.0569413 |
| Epi 1h | 31.6199 | 3.55868 | 0.559933 | 0.0641853 |
| SA 1h | 531.528 | 202.488 | 6.90589 | 2.34153 |
| Me-JA 1h | 281.877 | 59.5616 | 5.09748 | 0.61307 |
| Control 6h | 77.8758 | 22.0757 | 1.11284 | 0.276009 |
| ABA 6h | 92.3648 | 13.2281 | 1.30807 | 0.137077 |
| ACC 6h | 39.2641 | 10.2673 | 0.482769 | 0.116504 |
| BABA 6h | 51.5608 | 13.9839 | 0.646427 | 0.246684 |
| Chitin 6h | 55.7601 | 10.7425 | 0.778141 | 0.124338 |
| Epi 6h | 67.5286 | 12.1919 | 0.892749 | 0.137591 |
| SA 6h | 1047.67 | 365.46 | 13.4491 | 5.50369 |
| Me-JA 6h | 168.487 | 50.1052 | 2.36493 | 0.979461 |
Source Transcript PGSC0003DMT400017286 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G15550.1 | +2 | 1e-107 | 340 | 190/410 (46%) | indole-3-acetate beta-D-glucosyltransferase | chr4:8877877-8879301 REVERSE LENGTH=474 |
