Probe CUST_5230_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_5230_PI426222305 | JHI_St_60k_v1 | DMT400009112 | CGACTTGATTGTTAGCTGTCCATACCACTAGGTTCTTTTTAGAATGTTATGCAATATGTA |
All Microarray Probes Designed to Gene DMG400003541
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_4921_PI426222305 | JHI_St_60k_v1 | DMT400009111 | ATTGATCAGTATCAGAGTGCACTTGGAGTTGATATTTGGAGCACTCACTACGAGAGAACA |
| CUST_5230_PI426222305 | JHI_St_60k_v1 | DMT400009112 | CGACTTGATTGTTAGCTGTCCATACCACTAGGTTCTTTTTAGAATGTTATGCAATATGTA |
Microarray Signals from CUST_5230_PI426222305
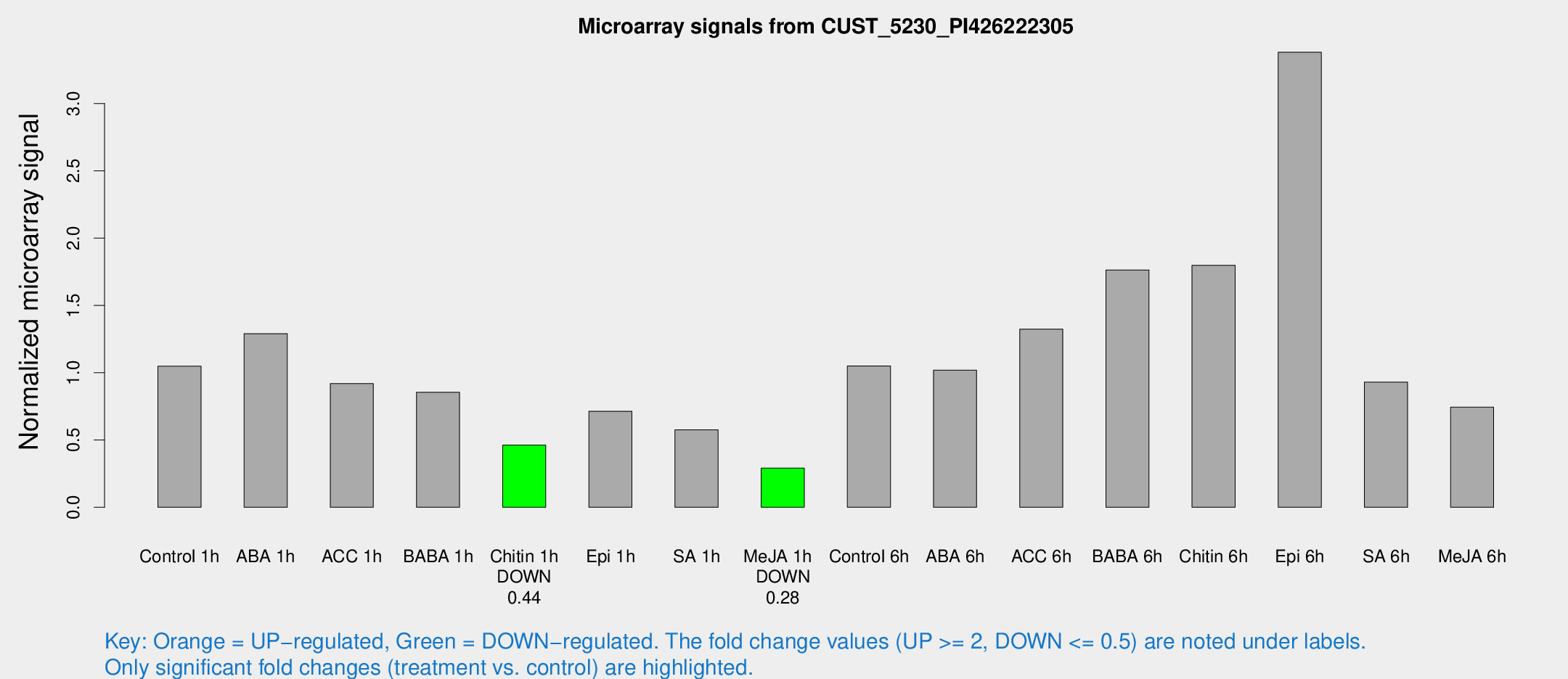
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 46.9554 | 8.42531 | 1.04831 | 0.121527 |
| ABA 1h | 51.0351 | 8.43601 | 1.28988 | 0.322245 |
| ACC 1h | 44.3031 | 10.958 | 0.918919 | 0.205894 |
| BABA 1h | 36.5215 | 4.72778 | 0.855196 | 0.187822 |
| Chitin 1h | 19.1378 | 4.28494 | 0.462756 | 0.0997683 |
| Epi 1h | 28.5867 | 6.69953 | 0.713923 | 0.174367 |
| SA 1h | 26.0544 | 3.99397 | 0.575414 | 0.0908974 |
| Me-JA 1h | 10.5561 | 3.8354 | 0.291257 | 0.109818 |
| Control 6h | 62.4094 | 25.218 | 1.04966 | 0.732784 |
| ABA 6h | 47.9335 | 6.11554 | 1.01906 | 0.184615 |
| ACC 6h | 73.0829 | 24.1769 | 1.32428 | 0.254858 |
| BABA 6h | 91.2498 | 20.1782 | 1.76358 | 0.404274 |
| Chitin 6h | 83.9395 | 7.63043 | 1.79887 | 0.189306 |
| Epi 6h | 174.24 | 39.8495 | 3.3818 | 0.869529 |
| SA 6h | 40.9198 | 6.3545 | 0.929903 | 0.111063 |
| Me-JA 6h | 35.0246 | 10.0682 | 0.743691 | 0.158819 |
Source Transcript PGSC0003DMT400009112 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G54340.1 | +3 | 1e-68 | 220 | 106/216 (49%) | K-box region and MADS-box transcription factor family protein | chr3:20119428-20121087 REVERSE LENGTH=232 |
