Probe CUST_52140_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_52140_PI426222305 | JHI_St_60k_v1 | DMT400014232 | AGCTTATTGTGAGTGTTATCTTGTTCGTATATGCATATTCTTTGCCTTGGCCAATTCAAG |
All Microarray Probes Designed to Gene DMG400005580
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_52140_PI426222305 | JHI_St_60k_v1 | DMT400014232 | AGCTTATTGTGAGTGTTATCTTGTTCGTATATGCATATTCTTTGCCTTGGCCAATTCAAG |
Microarray Signals from CUST_52140_PI426222305
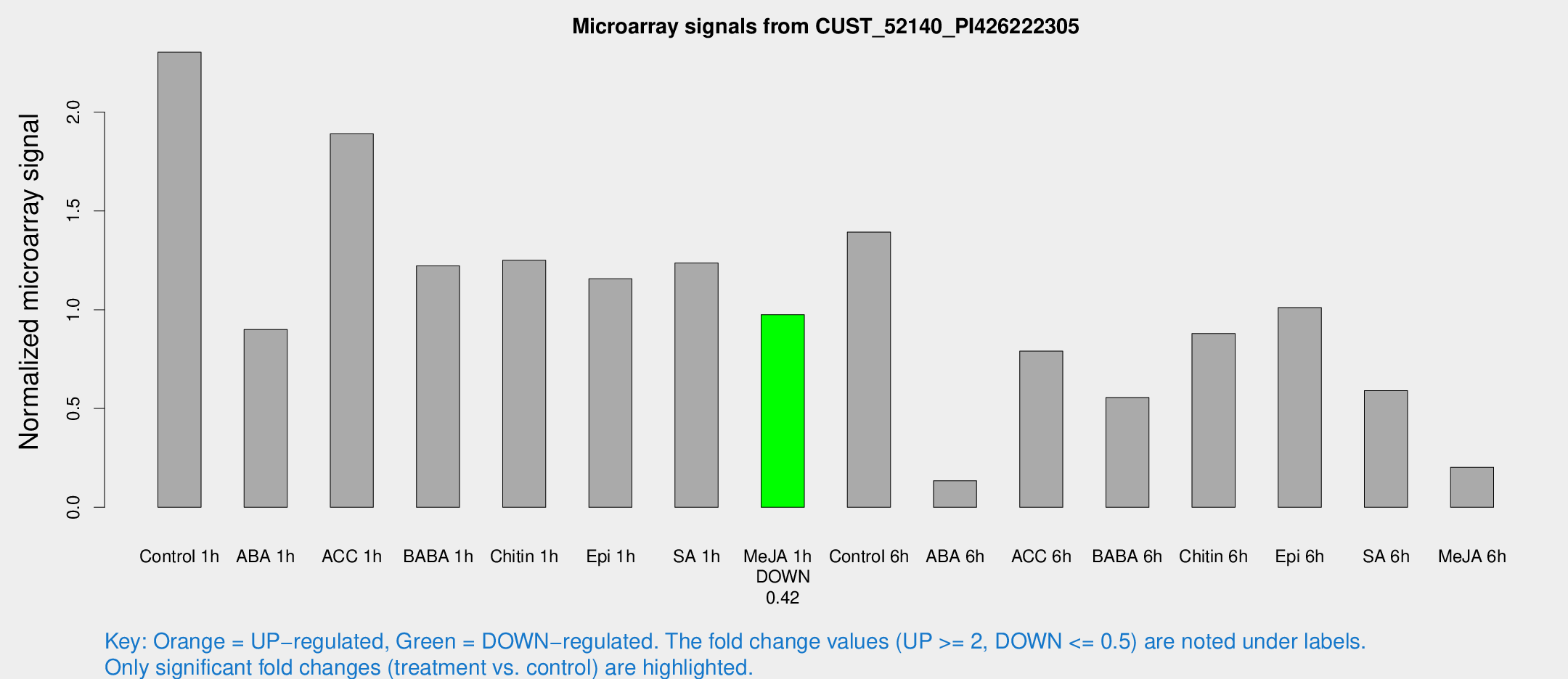
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 309.997 | 26.7049 | 2.30268 | 0.135314 |
| ABA 1h | 112.053 | 22.9994 | 0.899978 | 0.229333 |
| ACC 1h | 302.622 | 111.561 | 1.88958 | 0.680876 |
| BABA 1h | 157.722 | 9.78644 | 1.22201 | 0.117374 |
| Chitin 1h | 156.014 | 28.9525 | 1.24988 | 0.202394 |
| Epi 1h | 135.208 | 13.5404 | 1.15622 | 0.11052 |
| SA 1h | 184.255 | 51.6219 | 1.2364 | 0.43386 |
| Me-JA 1h | 107.294 | 10.1942 | 0.974692 | 0.144573 |
| Control 6h | 214.882 | 69.1113 | 1.39262 | 0.461546 |
| ABA 6h | 22.4966 | 9.54895 | 0.134314 | 0.0679778 |
| ACC 6h | 123.308 | 15.9314 | 0.790049 | 0.0605868 |
| BABA 6h | 84.6846 | 11.5505 | 0.555064 | 0.09238 |
| Chitin 6h | 126.302 | 12.4473 | 0.879093 | 0.105276 |
| Epi 6h | 165.671 | 48.0202 | 1.01023 | 0.41254 |
| SA 6h | 79.6272 | 10.5888 | 0.590459 | 0.140591 |
| Me-JA 6h | 61.371 | 47.6548 | 0.20255 | 0.355447 |
Source Transcript PGSC0003DMT400014232 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G07990.1 | +3 | 1e-114 | 355 | 194/505 (38%) | Cytochrome P450 superfamily protein | chr5:2560437-2562859 FORWARD LENGTH=513 |
