Probe CUST_52022_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_52022_PI426222305 | JHI_St_60k_v1 | DMT400059834 | GGAGATTCCAGAGAGAATGAAAGACGTGGAGCTGTTGAAGAGGAATTATAATAAGTGGTG |
All Microarray Probes Designed to Gene DMG400023274
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_52022_PI426222305 | JHI_St_60k_v1 | DMT400059834 | GGAGATTCCAGAGAGAATGAAAGACGTGGAGCTGTTGAAGAGGAATTATAATAAGTGGTG |
| CUST_52019_PI426222305 | JHI_St_60k_v1 | DMT400059836 | GGCTGTCAGTGAGAGTGGTTTTAGTGTAATTGGTTTGAACATAATTGTTTTTGCCAAAAA |
Microarray Signals from CUST_52022_PI426222305
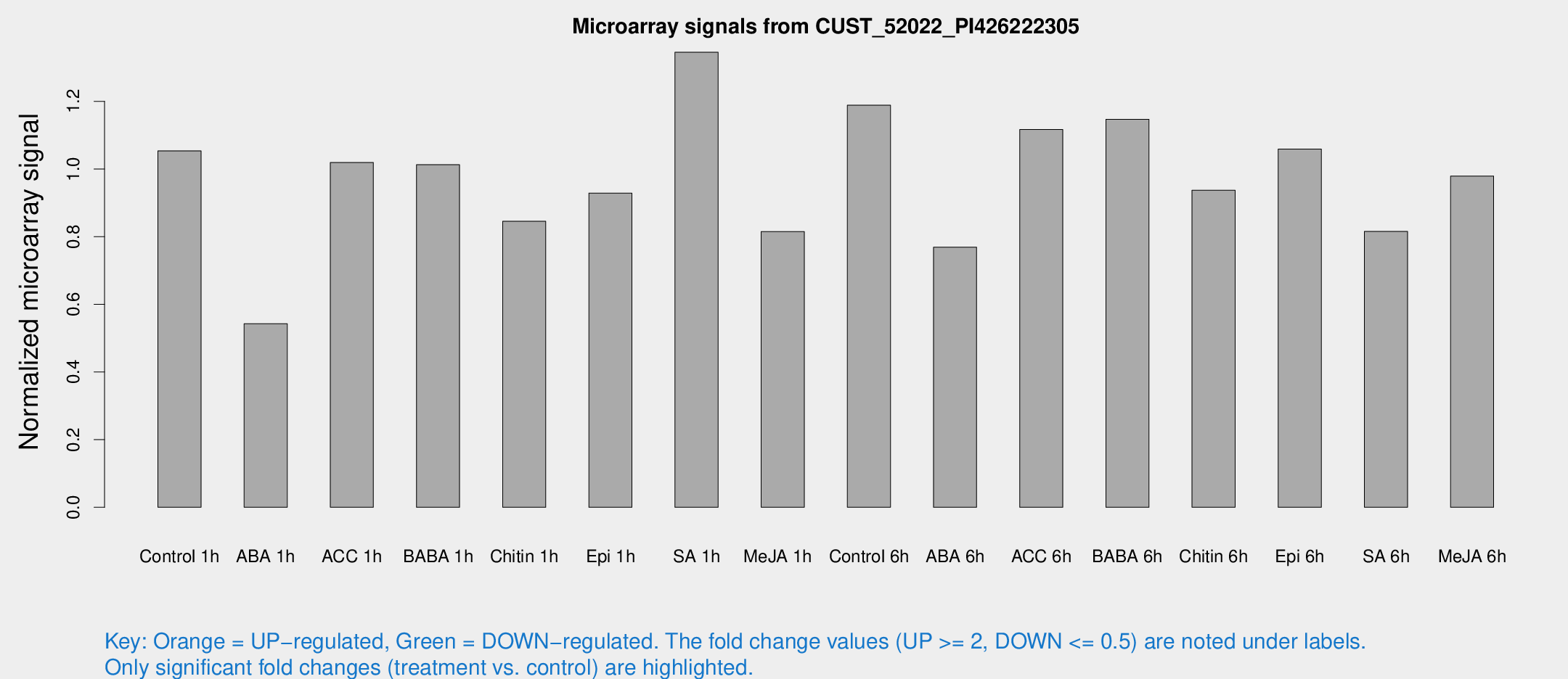
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 1387.49 | 202.082 | 1.05364 | 0.101393 |
| ABA 1h | 627.415 | 70.2607 | 0.542814 | 0.0314951 |
| ACC 1h | 1505.65 | 437.215 | 1.0191 | 0.29727 |
| BABA 1h | 1301.85 | 218.378 | 1.0127 | 0.082361 |
| Chitin 1h | 989.243 | 85.2239 | 0.84546 | 0.0489115 |
| Epi 1h | 1052.69 | 129.932 | 0.928761 | 0.0857329 |
| SA 1h | 1812.46 | 219.574 | 1.34511 | 0.122019 |
| Me-JA 1h | 876.009 | 126.644 | 0.815083 | 0.0527326 |
| Control 6h | 1647.09 | 384.336 | 1.18833 | 0.219699 |
| ABA 6h | 1078.65 | 165.02 | 0.768974 | 0.0732413 |
| ACC 6h | 1672.61 | 177.697 | 1.11657 | 0.0649383 |
| BABA 6h | 1682.13 | 205.949 | 1.14701 | 0.0959005 |
| Chitin 6h | 1298.94 | 115.742 | 0.937301 | 0.0608839 |
| Epi 6h | 1546.08 | 89.6715 | 1.0587 | 0.0612096 |
| SA 6h | 1171.01 | 342.195 | 0.815254 | 0.213336 |
| Me-JA 6h | 1284.59 | 173.989 | 0.979089 | 0.0664128 |
Source Transcript PGSC0003DMT400059834 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G59218.1 | +1 | 2e-19 | 90 | 84/284 (30%) | Disease resistance protein (CC-NBS-LRR class) family | chr1:21828398-21831713 FORWARD LENGTH=1049 |
