Probe CUST_51718_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_51718_PI426222305 | JHI_St_60k_v1 | DMT400080724 | GAGCTTATTGTATGTCCTCAATGCAATCACGAACCAAAATGATTCAATGAATACACAAAC |
All Microarray Probes Designed to Gene DMG400031437
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_51718_PI426222305 | JHI_St_60k_v1 | DMT400080724 | GAGCTTATTGTATGTCCTCAATGCAATCACGAACCAAAATGATTCAATGAATACACAAAC |
| CUST_51719_PI426222305 | JHI_St_60k_v1 | DMT400080723 | GAGCTTATTGTATGTCCTCAATGCAATCACGAACCAAAATGATTCAATGAATACACAAAC |
Microarray Signals from CUST_51718_PI426222305
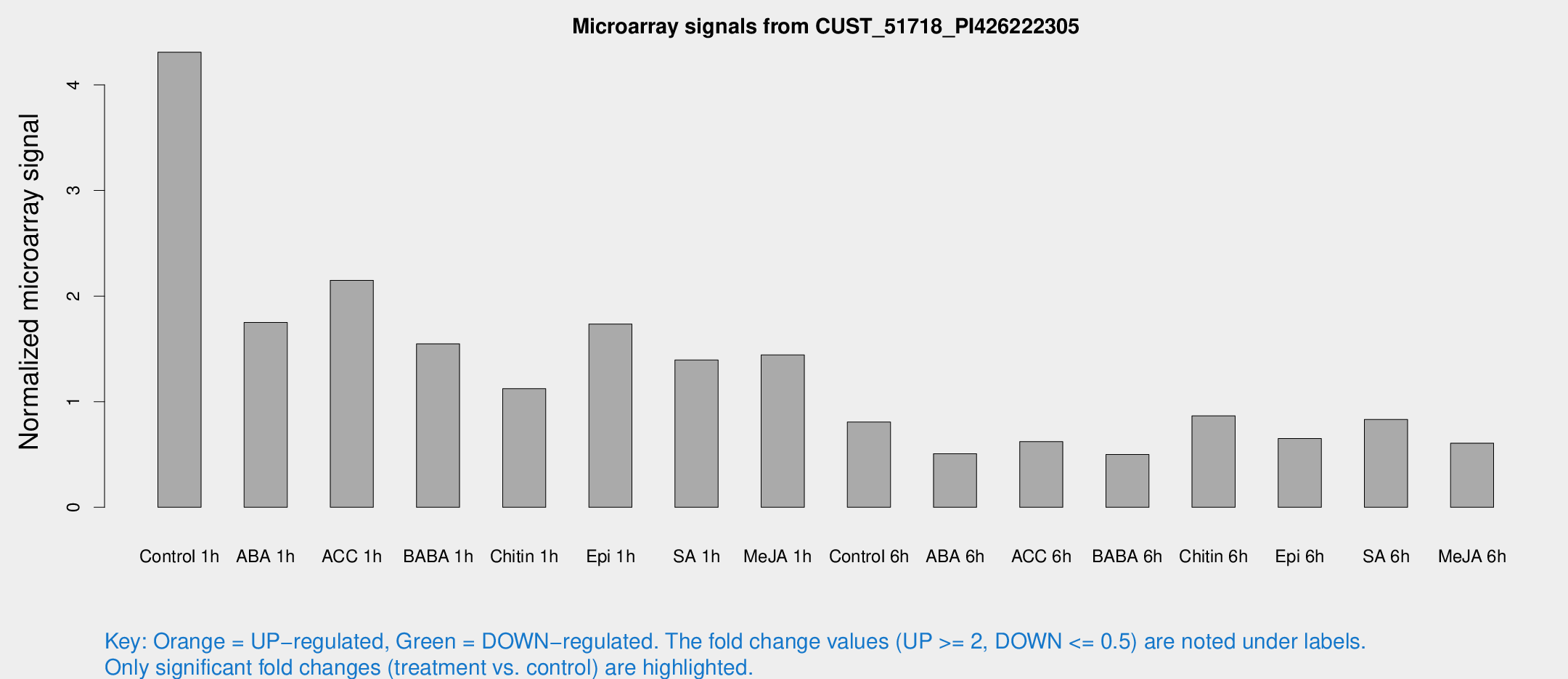
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 59.1372 | 18.8714 | 4.30913 | 1.05098 |
| ABA 1h | 20.9662 | 5.72715 | 1.75056 | 0.390138 |
| ACC 1h | 32.2875 | 10.6736 | 2.14968 | 0.824862 |
| BABA 1h | 19.4439 | 4.0855 | 1.54783 | 0.34893 |
| Chitin 1h | 15.6267 | 7.03515 | 1.12285 | 0.486537 |
| Epi 1h | 21.2096 | 7.62349 | 1.73552 | 0.652235 |
| SA 1h | 18.5088 | 3.69658 | 1.39435 | 0.353468 |
| Me-JA 1h | 16.4805 | 4.65824 | 1.44303 | 0.598482 |
| Control 6h | 12.6511 | 6.21874 | 0.807346 | 0.400792 |
| ABA 6h | 6.75811 | 3.81528 | 0.507676 | 0.285355 |
| ACC 6h | 9.12605 | 4.56629 | 0.622098 | 0.314846 |
| BABA 6h | 7.02996 | 4.0999 | 0.499938 | 0.290398 |
| Chitin 6h | 15.1435 | 8.15284 | 0.86641 | 0.489774 |
| Epi 6h | 9.42465 | 4.58628 | 0.651127 | 0.321949 |
| SA 6h | 10.7992 | 4.08881 | 0.831796 | 0.346215 |
| Me-JA 6h | 7.66021 | 3.73109 | 0.60702 | 0.301277 |
Source Transcript PGSC0003DMT400080724 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G58784.1 | +1 | 5e-54 | 146 | 75/131 (57%) | Undecaprenyl pyrophosphate synthetase family protein | chr5:23739950-23741267 REVERSE LENGTH=302 |
