Probe CUST_51648_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_51648_PI426222305 | JHI_St_60k_v1 | DMT400011402 | GGCTTTGCCAAATGATGAAAGGAAGTCATATAAATATACTTGGACTCTTCCAAGAATATG |
All Microarray Probes Designed to Gene DMG400004467
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_51648_PI426222305 | JHI_St_60k_v1 | DMT400011402 | GGCTTTGCCAAATGATGAAAGGAAGTCATATAAATATACTTGGACTCTTCCAAGAATATG |
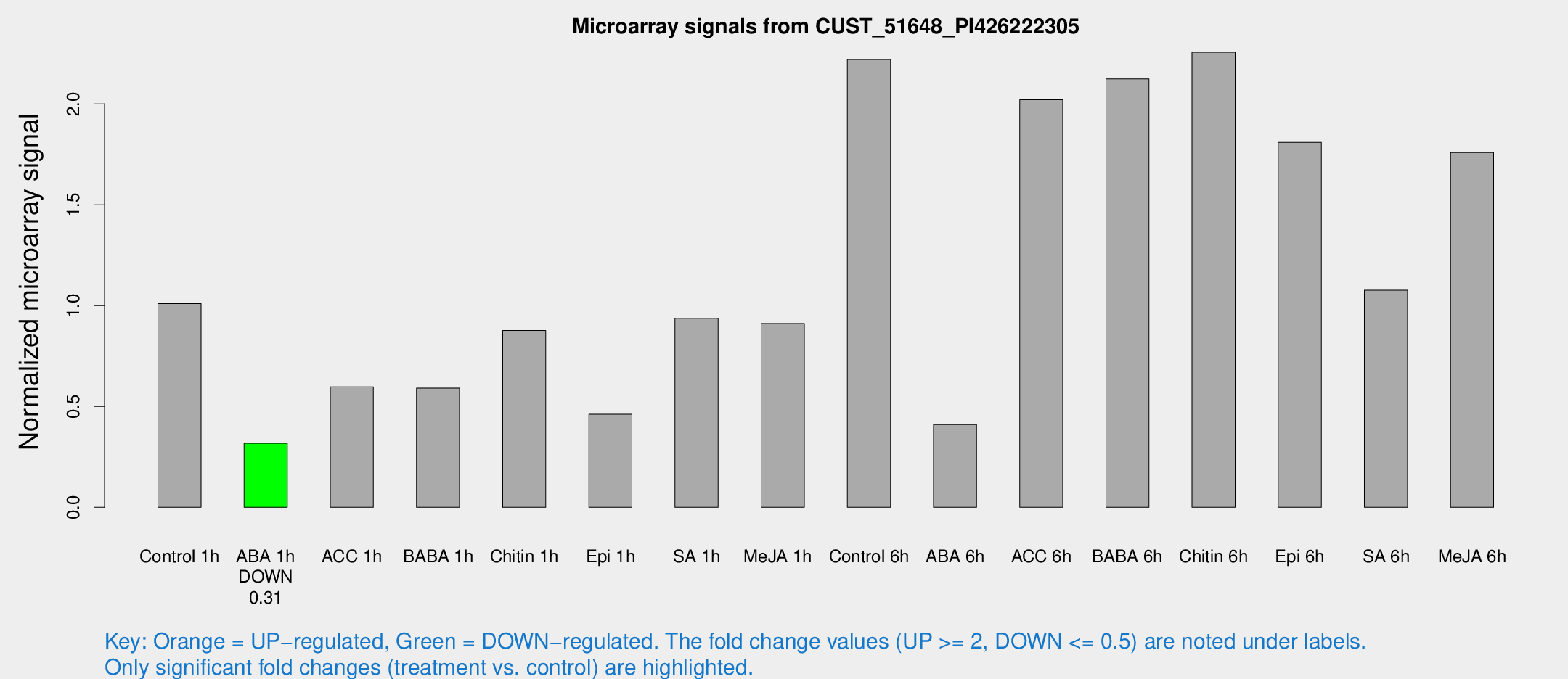
Microarray Signals from CUST_51648_PI426222305
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 22.0797 | 3.57483 | 1.00993 | 0.169431 |
| ABA 1h | 6.09586 | 3.23386 | 0.317303 | 0.171146 |
| ACC 1h | 14.5183 | 4.7983 | 0.596864 | 0.227243 |
| BABA 1h | 13.6344 | 3.8126 | 0.591125 | 0.283564 |
| Chitin 1h | 16.9211 | 3.45422 | 0.876486 | 0.179248 |
| Epi 1h | 10.0255 | 4.16947 | 0.461323 | 0.212519 |
| SA 1h | 22.6924 | 6.79707 | 0.937212 | 0.387357 |
| Me-JA 1h | 16.8977 | 4.11796 | 0.911377 | 0.26417 |
| Control 6h | 54.4256 | 19.9437 | 2.21969 | 0.71869 |
| ABA 6h | 9.86374 | 3.60284 | 0.410145 | 0.167554 |
| ACC 6h | 51.6727 | 9.81629 | 2.02015 | 0.277486 |
| BABA 6h | 59.432 | 19.366 | 2.12399 | 0.915166 |
| Chitin 6h | 51.9682 | 4.88273 | 2.25603 | 0.214288 |
| Epi 6h | 45.2233 | 7.54731 | 1.80964 | 0.484996 |
| SA 6h | 27.3696 | 9.41545 | 1.07618 | 0.702249 |
| Me-JA 6h | 43.2863 | 13.4346 | 1.75867 | 0.64348 |
Source Transcript PGSC0003DMT400011402 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G14590.1 | +1 | 4e-124 | 367 | 161/270 (60%) | Nucleotide-diphospho-sugar transferase family protein | chr1:4998957-5000617 REVERSE LENGTH=386 |
