Probe CUST_51620_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_51620_PI426222305 | JHI_St_60k_v1 | DMT400053852 | TAAACTACAGAGCGAAGAGATTGCAAACACGATAAGGAAGGTGTTAGTTGAGGAAAGTGG |
All Microarray Probes Designed to Gene DMG400020895
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_51620_PI426222305 | JHI_St_60k_v1 | DMT400053852 | TAAACTACAGAGCGAAGAGATTGCAAACACGATAAGGAAGGTGTTAGTTGAGGAAAGTGG |
| CUST_51625_PI426222305 | JHI_St_60k_v1 | DMT400053853 | AGAATGCTCTTTGGCGCGAAGAGGGAAGAAAGTTGAGGTTACTGTAAATTCCAAAATCTG |
Microarray Signals from CUST_51620_PI426222305
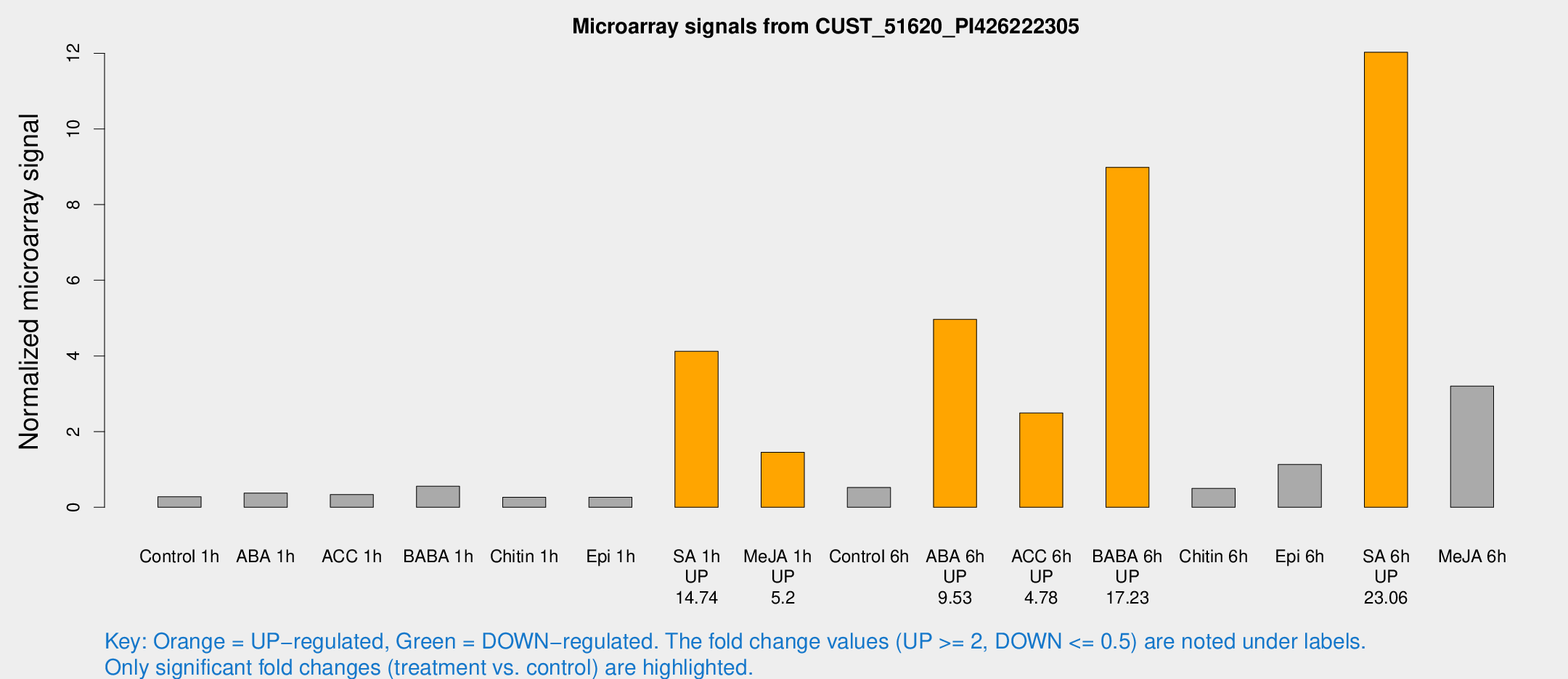
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 6.95657 | 3.32859 | 0.279592 | 0.141459 |
| ABA 1h | 9.32548 | 3.70406 | 0.377012 | 0.186582 |
| ACC 1h | 9.1138 | 3.52565 | 0.33854 | 0.151942 |
| BABA 1h | 16.3151 | 7.47226 | 0.556952 | 0.426978 |
| Chitin 1h | 5.83009 | 3.31832 | 0.264727 | 0.150249 |
| Epi 1h | 5.60601 | 3.25019 | 0.267001 | 0.154612 |
| SA 1h | 110.279 | 26.3539 | 4.12169 | 0.972953 |
| Me-JA 1h | 33.2519 | 10.6237 | 1.45428 | 0.558908 |
| Control 6h | 14.6512 | 5.48204 | 0.521598 | 0.18999 |
| ABA 6h | 174.107 | 92.7007 | 4.96844 | 3.09729 |
| ACC 6h | 70.0767 | 7.71632 | 2.49244 | 0.585234 |
| BABA 6h | 258.102 | 56.917 | 8.98474 | 2.5565 |
| Chitin 6h | 13.5089 | 4.1915 | 0.497599 | 0.17721 |
| Epi 6h | 31.1278 | 4.51824 | 1.13401 | 0.162178 |
| SA 6h | 299.46 | 59.5035 | 12.0267 | 1.5591 |
| Me-JA 6h | 107.77 | 55.1396 | 3.20438 | 3.06763 |
Source Transcript PGSC0003DMT400053852 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G65550.1 | +1 | 4e-28 | 107 | 55/119 (46%) | UDP-Glycosyltransferase superfamily protein | chr5:26198410-26199810 REVERSE LENGTH=466 |
