Probe CUST_51594_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_51594_PI426222305 | JHI_St_60k_v1 | DMT400005537 | CTGCTGCAACACCTCCTTATTCGAATTGATTTCTCGTATGATCACTTGATTAATAGTTAT |
All Microarray Probes Designed to Gene DMG402002164
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_51592_PI426222305 | JHI_St_60k_v1 | DMT400005536 | CGTTTTTGGGTTAAGTTTTTCGAAGATAAGGGGTCAGCAGTTGGGACAAGTTTCTTTCAC |
| CUST_51594_PI426222305 | JHI_St_60k_v1 | DMT400005537 | CTGCTGCAACACCTCCTTATTCGAATTGATTTCTCGTATGATCACTTGATTAATAGTTAT |
Microarray Signals from CUST_51594_PI426222305
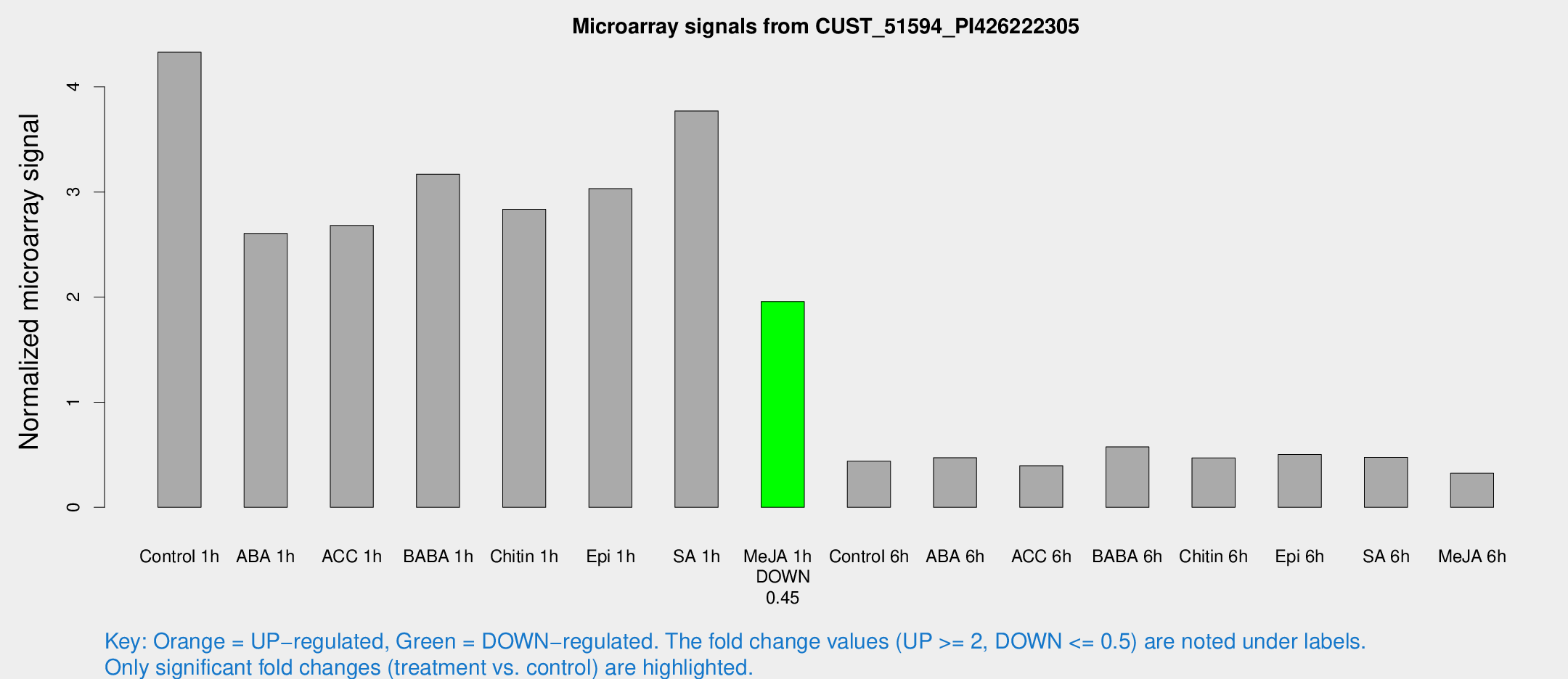
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 10242.2 | 897.304 | 4.32982 | 0.249986 |
| ABA 1h | 5545.37 | 874.129 | 2.60674 | 0.22204 |
| ACC 1h | 7736.48 | 2695.84 | 2.68193 | 1.15509 |
| BABA 1h | 7556.08 | 1571.5 | 3.16907 | 0.428843 |
| Chitin 1h | 5988.76 | 346.012 | 2.83509 | 0.16369 |
| Epi 1h | 6161.26 | 355.75 | 3.03223 | 0.175072 |
| SA 1h | 9208.71 | 982.706 | 3.77145 | 0.243764 |
| Me-JA 1h | 3815.88 | 487.924 | 1.95792 | 0.113053 |
| Control 6h | 1067.31 | 183.181 | 0.438936 | 0.0463307 |
| ABA 6h | 1220.96 | 223.536 | 0.472127 | 0.0654062 |
| ACC 6h | 1069.51 | 72.4552 | 0.395368 | 0.0323877 |
| BABA 6h | 1526.98 | 160.824 | 0.57438 | 0.0393499 |
| Chitin 6h | 1188.89 | 134.594 | 0.469794 | 0.0451215 |
| Epi 6h | 1351.1 | 172.233 | 0.501823 | 0.047881 |
| SA 6h | 1226.38 | 386.654 | 0.475777 | 0.113309 |
| Me-JA 6h | 819.498 | 231.061 | 0.324432 | 0.0648668 |
Source Transcript PGSC0003DMT400005537 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G09270.1 | +1 | 3e-15 | 71 | 40/94 (43%) | glutathione S-transferase TAU 8 | chr3:2848407-2849226 REVERSE LENGTH=224 |
