Probe CUST_51403_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_51403_PI426222305 | JHI_St_60k_v1 | DMT400080543 | CTGATTATCCAACTCTAGAAACCATGACATGATTGGATTCCTTGGTAACTAATTATCAGA |
All Microarray Probes Designed to Gene DMG400031362
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_51403_PI426222305 | JHI_St_60k_v1 | DMT400080543 | CTGATTATCCAACTCTAGAAACCATGACATGATTGGATTCCTTGGTAACTAATTATCAGA |
| CUST_51401_PI426222305 | JHI_St_60k_v1 | DMT400080542 | TGAGGAAGGCATATAAAGATTTACCAAAGCGATTTGAGCTAAATGCTCCTCTTAACAAGT |
Microarray Signals from CUST_51403_PI426222305
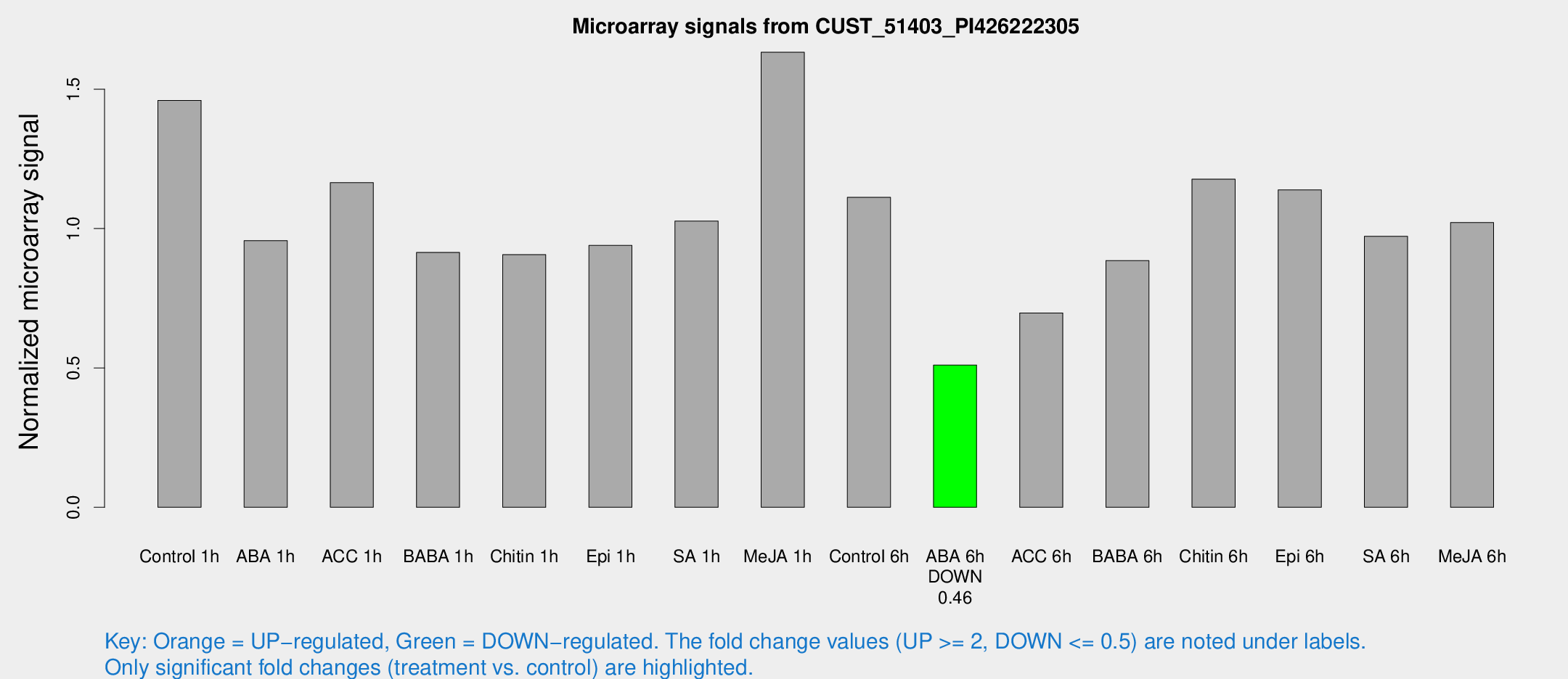
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 3392.55 | 491.898 | 1.45992 | 0.121238 |
| ABA 1h | 1943.93 | 171.408 | 0.956752 | 0.0552628 |
| ACC 1h | 2816.01 | 504.803 | 1.16457 | 0.149265 |
| BABA 1h | 2103.69 | 413.282 | 0.914476 | 0.109306 |
| Chitin 1h | 1874.01 | 172.872 | 0.906301 | 0.0573117 |
| Epi 1h | 1862.12 | 129.946 | 0.939497 | 0.0658452 |
| SA 1h | 2476.74 | 406.94 | 1.02676 | 0.144301 |
| Me-JA 1h | 3052.99 | 216.936 | 1.63267 | 0.0942813 |
| Control 6h | 2624.41 | 442.409 | 1.11214 | 0.110125 |
| ABA 6h | 1240.56 | 71.9572 | 0.510417 | 0.0564265 |
| ACC 6h | 1851.04 | 224.4 | 0.69734 | 0.045887 |
| BABA 6h | 2312.36 | 368.986 | 0.885186 | 0.127546 |
| Chitin 6h | 2859.33 | 165.336 | 1.17729 | 0.0679908 |
| Epi 6h | 2959.07 | 320.869 | 1.13857 | 0.166535 |
| SA 6h | 2247.56 | 333.822 | 0.972372 | 0.0561691 |
| Me-JA 6h | 2416.21 | 505.942 | 1.02157 | 0.127798 |
Source Transcript PGSC0003DMT400080543 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G76690.1 | +3 | 1e-37 | 98 | 45/71 (63%) | 12-oxophytodienoate reductase 2 | chr1:28778976-28780355 FORWARD LENGTH=374 |
