Probe CUST_51402_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_51402_PI426222305 | JHI_St_60k_v1 | DMT400080548 | GCTCCACTTCCAATCTGTTAGAAGTGAAGCAACAGGTTTTATTATGTTAATGTTTGTCAA |
All Microarray Probes Designed to Gene DMG400031365
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_51402_PI426222305 | JHI_St_60k_v1 | DMT400080548 | GCTCCACTTCCAATCTGTTAGAAGTGAAGCAACAGGTTTTATTATGTTAATGTTTGTCAA |
Microarray Signals from CUST_51402_PI426222305
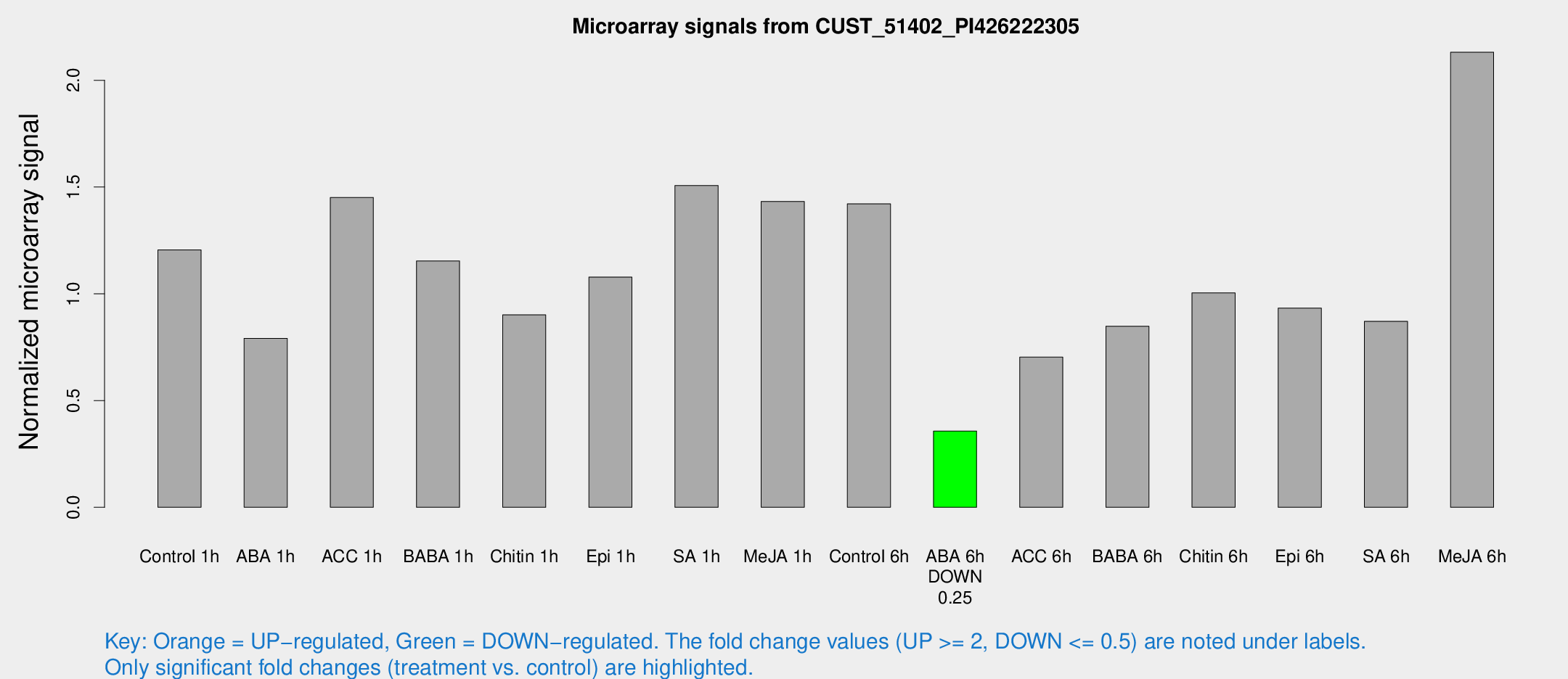
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 2612.16 | 710.537 | 1.20529 | 0.278804 |
| ABA 1h | 1436.88 | 176.11 | 0.791425 | 0.102871 |
| ACC 1h | 3268.27 | 849.482 | 1.45128 | 0.344419 |
| BABA 1h | 2258.76 | 166.43 | 1.15414 | 0.0689921 |
| Chitin 1h | 1647.88 | 133.959 | 0.901327 | 0.062943 |
| Epi 1h | 1892.45 | 118.461 | 1.07831 | 0.0622818 |
| SA 1h | 3435.61 | 1095.47 | 1.507 | 0.51865 |
| Me-JA 1h | 2367.26 | 137.029 | 1.43245 | 0.082726 |
| Control 6h | 3266.04 | 1178.35 | 1.42106 | 0.412594 |
| ABA 6h | 788.547 | 123.008 | 0.357171 | 0.0666203 |
| ACC 6h | 1671.23 | 254.357 | 0.703287 | 0.0979168 |
| BABA 6h | 1928.71 | 150.135 | 0.848155 | 0.0784669 |
| Chitin 6h | 2197.44 | 296.988 | 1.00441 | 0.148527 |
| Epi 6h | 2418.67 | 731.709 | 0.933268 | 0.503683 |
| SA 6h | 1756.37 | 170.806 | 0.871189 | 0.0688118 |
| Me-JA 6h | 4384.14 | 650.422 | 2.13135 | 0.141403 |
Source Transcript PGSC0003DMT400080548 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G37040.1 | +2 | 0.0 | 1140 | 574/694 (83%) | PHE ammonia lyase 1 | chr2:15557602-15560237 REVERSE LENGTH=725 |
