Probe CUST_51201_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_51201_PI426222305 | JHI_St_60k_v1 | DMT400065999 | CTATGTTCATTGGTTATGGTTCAATACCATATTGACCCAATGGATAAGCTATAAAGTGCT |
All Microarray Probes Designed to Gene DMG400025692
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_51200_PI426222305 | JHI_St_60k_v1 | DMT400066000 | CTATGTTCATTGGTTATGGTTCAATACCATATTGACCCAATGGATAAGCTATAAAGTGCT |
| CUST_51201_PI426222305 | JHI_St_60k_v1 | DMT400065999 | CTATGTTCATTGGTTATGGTTCAATACCATATTGACCCAATGGATAAGCTATAAAGTGCT |
Microarray Signals from CUST_51201_PI426222305
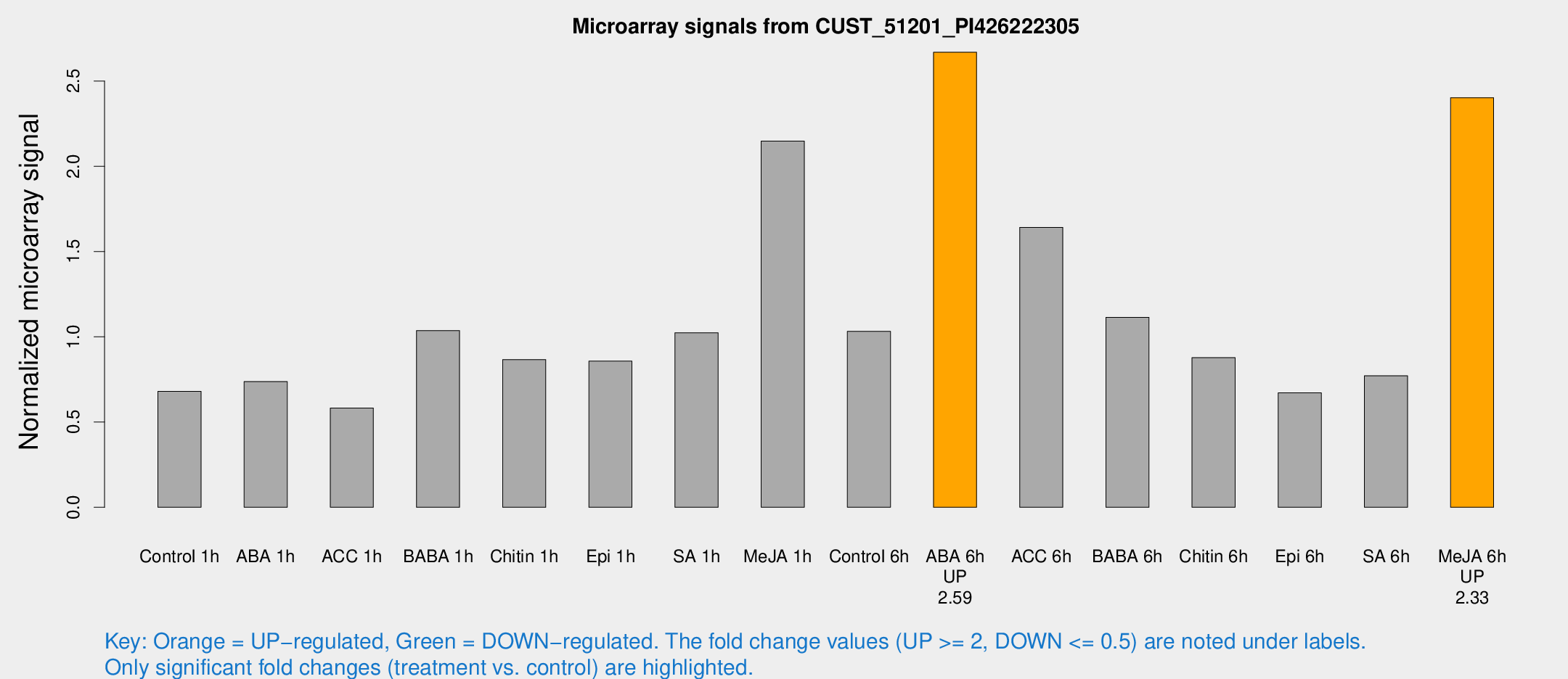
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 206.904 | 17.764 | 0.680081 | 0.0935481 |
| ABA 1h | 221.878 | 66.8371 | 0.736943 | 0.240818 |
| ACC 1h | 238.019 | 113.37 | 0.582221 | 0.34332 |
| BABA 1h | 313.284 | 55.6149 | 1.03683 | 0.0981721 |
| Chitin 1h | 257.716 | 73.7864 | 0.866367 | 0.263744 |
| Epi 1h | 235.855 | 50.0097 | 0.857139 | 0.200082 |
| SA 1h | 322.555 | 38.7351 | 1.02314 | 0.0855969 |
| Me-JA 1h | 544.489 | 90.5566 | 2.14781 | 0.171133 |
| Control 6h | 321.726 | 51.2428 | 1.03238 | 0.0819266 |
| ABA 6h | 902.743 | 186.143 | 2.66901 | 0.54246 |
| ACC 6h | 571.655 | 39.769 | 1.64204 | 0.296191 |
| BABA 6h | 399.563 | 101.662 | 1.11439 | 0.239729 |
| Chitin 6h | 282.436 | 16.8879 | 0.877522 | 0.0524252 |
| Epi 6h | 231.037 | 21.8668 | 0.671932 | 0.040911 |
| SA 6h | 236.06 | 38.2416 | 0.770993 | 0.097386 |
| Me-JA 6h | 730.589 | 77.9476 | 2.40237 | 0.139329 |
Source Transcript PGSC0003DMT400065999 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G69040.2 | +2 | 0.0 | 654 | 335/454 (74%) | ACT domain repeat 4 | chr1:25957843-25960079 FORWARD LENGTH=455 |
