Probe CUST_50931_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_50931_PI426222305 | JHI_St_60k_v1 | DMT400013962 | AATAGTGATAAGGTCCGGATACATTCTAACCTACCCAGACTCCACTTAATGAAATTTCAC |
All Microarray Probes Designed to Gene DMG400005468
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_50925_PI426222305 | JHI_St_60k_v1 | DMT400013961 | GGTGCATTTGGCAAGGAAAGTGTAAACGATCTAAGAAAGTTTTATGGACTTATAAGGATA |
| CUST_50931_PI426222305 | JHI_St_60k_v1 | DMT400013962 | AATAGTGATAAGGTCCGGATACATTCTAACCTACCCAGACTCCACTTAATGAAATTTCAC |
Microarray Signals from CUST_50931_PI426222305
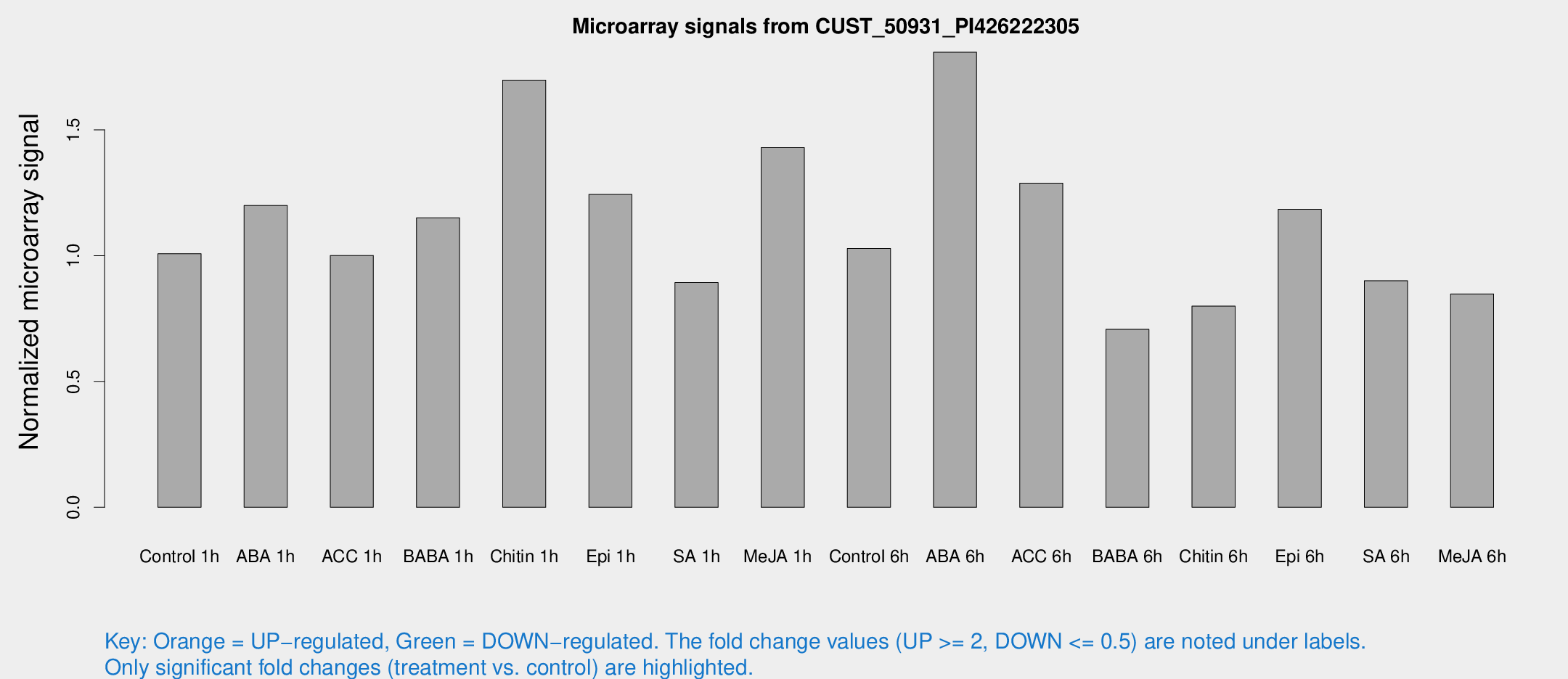
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 9.55062 | 4.67811 | 1.00772 | 0.449844 |
| ABA 1h | 8.91358 | 2.84887 | 1.19896 | 0.46556 |
| ACC 1h | 9.27505 | 3.7621 | 1.00049 | 0.465764 |
| BABA 1h | 9.21169 | 3.09719 | 1.15038 | 0.450497 |
| Chitin 1h | 13.0732 | 3.75785 | 1.69739 | 0.503818 |
| Epi 1h | 9.01716 | 2.93441 | 1.24312 | 0.487076 |
| SA 1h | 7.22633 | 2.9332 | 0.892755 | 0.371101 |
| Me-JA 1h | 9.53748 | 3.03101 | 1.42898 | 0.53389 |
| Control 6h | 9.8775 | 4.71537 | 1.0284 | 0.693323 |
| ABA 6h | 19.0686 | 7.53199 | 1.80811 | 1.13826 |
| ACC 6h | 11.5955 | 3.72986 | 1.28784 | 0.410366 |
| BABA 6h | 6.17091 | 3.41565 | 0.707274 | 0.391566 |
| Chitin 6h | 6.66036 | 3.35684 | 0.799543 | 0.407676 |
| Epi 6h | 12.2427 | 4.85141 | 1.18377 | 0.493387 |
| SA 6h | 7.33701 | 3.21724 | 0.900422 | 0.441826 |
| Me-JA 6h | 6.64047 | 3.05667 | 0.847614 | 0.400376 |
Source Transcript PGSC0003DMT400013962 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G46620.1 | +3 | 7e-162 | 479 | 271/493 (55%) | P-loop containing nucleoside triphosphate hydrolases superfamily protein | chr2:19139071-19140546 REVERSE LENGTH=491 |
