Probe CUST_50845_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_50845_PI426222305 | JHI_St_60k_v1 | DMT400080659 | CGTATTCATCTTTTCTTAATGTGCTCACACTTATGGCTCTTACACACCATCTTGTTCACC |
All Microarray Probes Designed to Gene DMG400031411
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_50845_PI426222305 | JHI_St_60k_v1 | DMT400080659 | CGTATTCATCTTTTCTTAATGTGCTCACACTTATGGCTCTTACACACCATCTTGTTCACC |
| CUST_50839_PI426222305 | JHI_St_60k_v1 | DMT400080660 | CCTATTTGGCTTCCCTCTATTTCGTCAAGTATACTAAGCTAGTTTTGATCATAGATTTTC |
Microarray Signals from CUST_50845_PI426222305
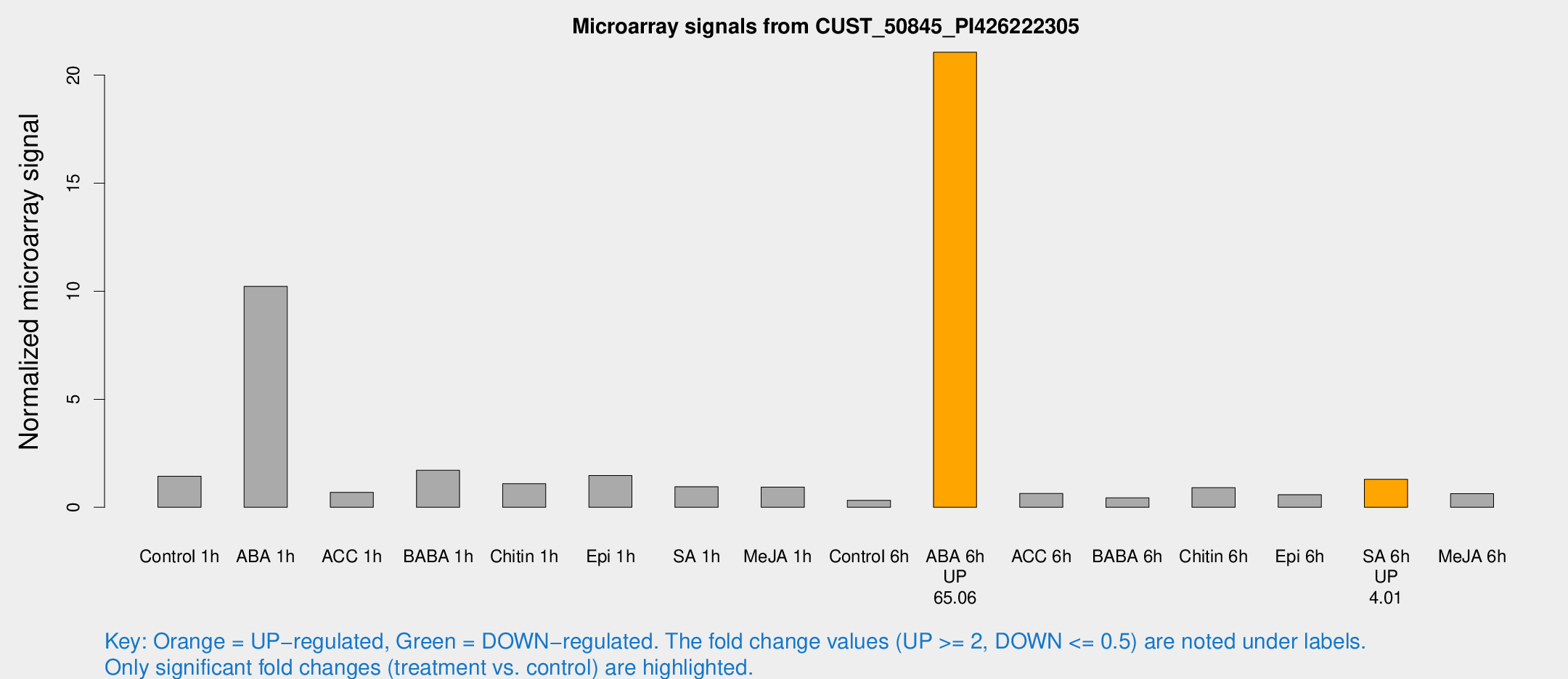
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 33.797 | 6.43895 | 1.43605 | 0.347944 |
| ABA 1h | 224.637 | 58.4201 | 10.2291 | 2.80684 |
| ACC 1h | 18.5636 | 5.66536 | 0.694811 | 0.243543 |
| BABA 1h | 40.6886 | 11.2238 | 1.71095 | 0.362106 |
| Chitin 1h | 26.9927 | 9.34505 | 1.09218 | 0.643029 |
| Epi 1h | 31.1066 | 8.68438 | 1.47301 | 0.415909 |
| SA 1h | 25.246 | 9.4669 | 0.953717 | 0.300568 |
| Me-JA 1h | 18.1014 | 3.79568 | 0.93701 | 0.182919 |
| Control 6h | 8.13766 | 3.16264 | 0.323745 | 0.153877 |
| ABA 6h | 515.115 | 58.3845 | 21.0615 | 1.98194 |
| ACC 6h | 18.3254 | 4.93334 | 0.638788 | 0.354961 |
| BABA 6h | 12.4975 | 4.06472 | 0.434953 | 0.160805 |
| Chitin 6h | 22.003 | 3.69037 | 0.909858 | 0.153136 |
| Epi 6h | 20.2329 | 10.2089 | 0.585327 | 0.353075 |
| SA 6h | 32.4357 | 10.8428 | 1.29763 | 0.450319 |
| Me-JA 6h | 15.1499 | 3.37509 | 0.630431 | 0.156554 |
Source Transcript PGSC0003DMT400080659 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G72100.1 | +1 | 3e-34 | 127 | 65/116 (56%) | late embryogenesis abundant domain-containing protein / LEA domain-containing protein | chr1:27127250-27128811 FORWARD LENGTH=480 |
