Probe CUST_50581_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_50581_PI426222305 | JHI_St_60k_v1 | DMT400068786 | TATGCAATGATCTCCCAATACTCAAAGAGCTTTTATCTATCCAGAAACAAGCTTGATGGT |
All Microarray Probes Designed to Gene DMG400026748
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_50581_PI426222305 | JHI_St_60k_v1 | DMT400068786 | TATGCAATGATCTCCCAATACTCAAAGAGCTTTTATCTATCCAGAAACAAGCTTGATGGT |
| CUST_50584_PI426222305 | JHI_St_60k_v1 | DMT400068782 | GTAGATGCCAATTTGATAACAGTACCAATGGATAATTGCTTACAGAAGGAGCTAGATGTT |
Microarray Signals from CUST_50581_PI426222305
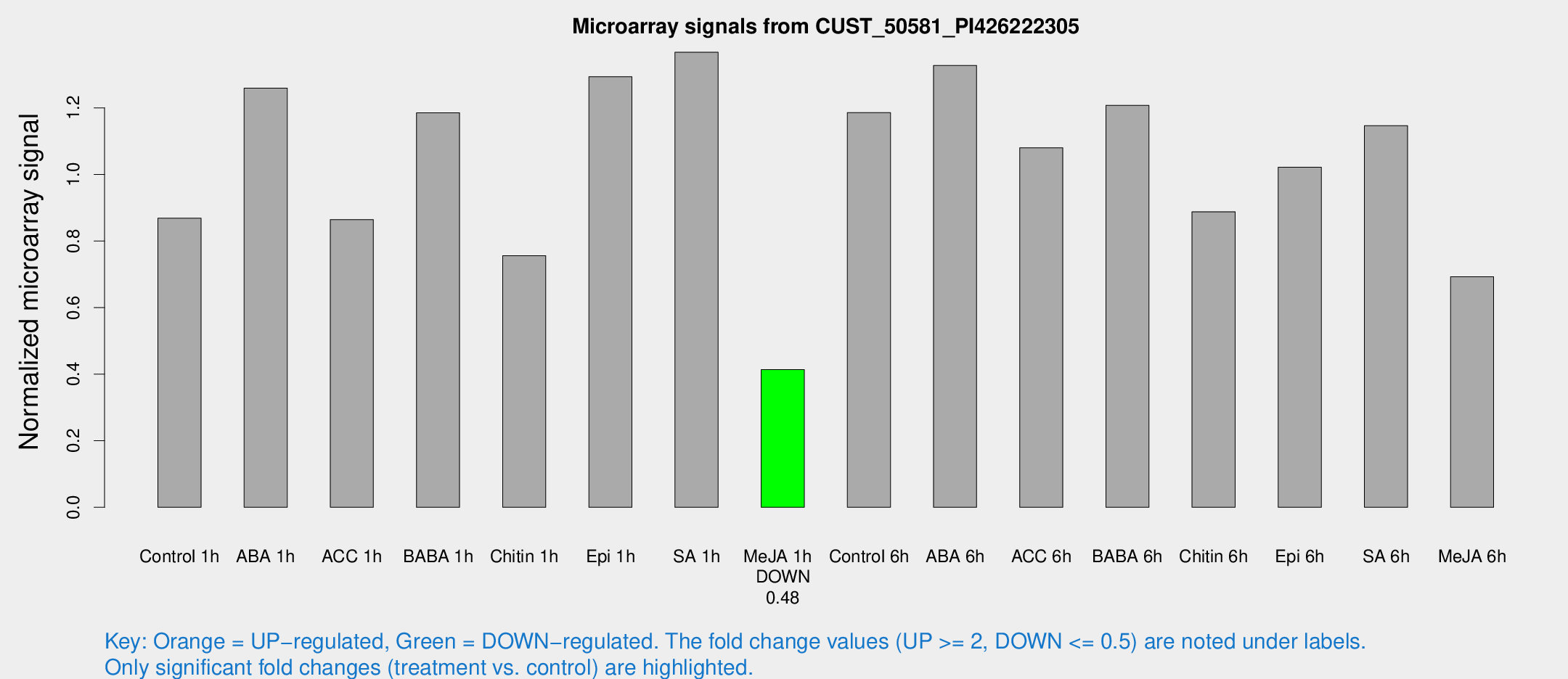
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 85.6207 | 8.72073 | 0.868381 | 0.06091 |
| ABA 1h | 116.62 | 32.2089 | 1.25938 | 0.248262 |
| ACC 1h | 90.2198 | 16.7383 | 0.864509 | 0.112174 |
| BABA 1h | 119.855 | 29.7086 | 1.18519 | 0.217386 |
| Chitin 1h | 66.5363 | 5.12171 | 0.756087 | 0.0579273 |
| Epi 1h | 123.469 | 38.9291 | 1.29376 | 0.489903 |
| SA 1h | 144.804 | 31.0104 | 1.36716 | 0.346336 |
| Me-JA 1h | 33.2787 | 3.98047 | 0.413623 | 0.0490902 |
| Control 6h | 121.724 | 25.4147 | 1.18579 | 0.17795 |
| ABA 6h | 141.447 | 22.1548 | 1.32746 | 0.150631 |
| ACC 6h | 125.895 | 23.1281 | 1.08027 | 0.192005 |
| BABA 6h | 137.946 | 26.7959 | 1.20764 | 0.221484 |
| Chitin 6h | 97.8608 | 24.383 | 0.887919 | 0.200716 |
| Epi 6h | 115.176 | 16.7486 | 1.02174 | 0.100969 |
| SA 6h | 112.211 | 12.6647 | 1.14655 | 0.264651 |
| Me-JA 6h | 77.2802 | 25.3343 | 0.692856 | 0.230489 |
Source Transcript PGSC0003DMT400068786 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G20480.1 | +1 | 7e-42 | 158 | 85/219 (39%) | EF-TU receptor | chr5:6922497-6925679 FORWARD LENGTH=1031 |
