Probe CUST_49703_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_49703_PI426222305 | JHI_St_60k_v1 | DMT400034372 | TGGTAACTTTTCTGAAGAAGAAGATGAACTCATTATCAAACTCCATAGCCTCCTTGGAAA |
All Microarray Probes Designed to Gene DMG400013215
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_49703_PI426222305 | JHI_St_60k_v1 | DMT400034372 | TGGTAACTTTTCTGAAGAAGAAGATGAACTCATTATCAAACTCCATAGCCTCCTTGGAAA |
| CUST_49704_PI426222305 | JHI_St_60k_v1 | DMT400034371 | TATGATAGGTGGTCGCTTATAGCAGGAAGATTACCAGGAAGAACAGATAACGAGATAAAA |
Microarray Signals from CUST_49703_PI426222305
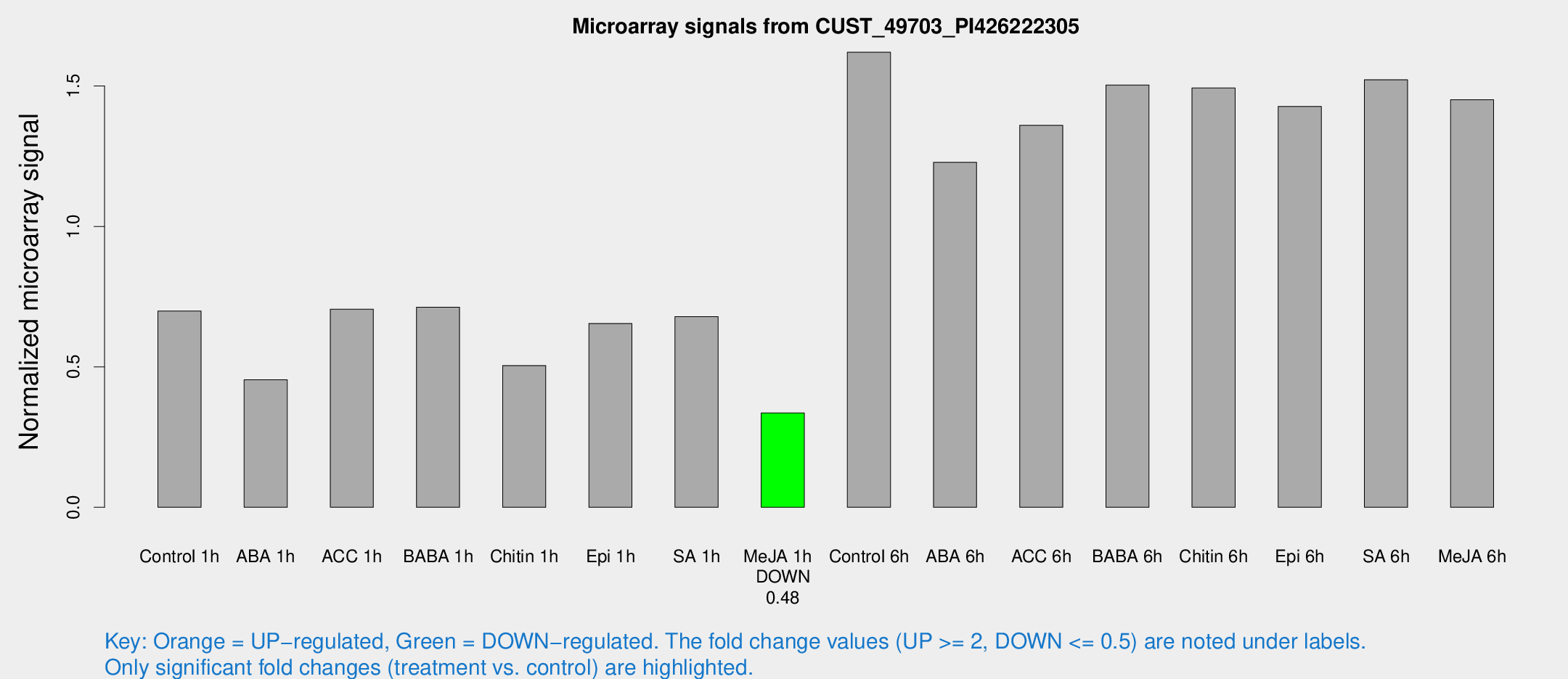
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 489.842 | 75.0859 | 0.698601 | 0.0674262 |
| ABA 1h | 280.354 | 34.5068 | 0.454367 | 0.0334039 |
| ACC 1h | 565.852 | 169.364 | 0.705441 | 0.237327 |
| BABA 1h | 483.347 | 72.2971 | 0.712544 | 0.0420386 |
| Chitin 1h | 317.739 | 46.7991 | 0.504766 | 0.0434937 |
| Epi 1h | 405.482 | 82.1258 | 0.654465 | 0.123648 |
| SA 1h | 482.392 | 41.9203 | 0.679113 | 0.0648168 |
| Me-JA 1h | 189.603 | 16.5507 | 0.336074 | 0.0203719 |
| Control 6h | 1193.48 | 284.435 | 1.6202 | 0.312189 |
| ABA 6h | 914.514 | 125.855 | 1.22838 | 0.116652 |
| ACC 6h | 1108.76 | 196.706 | 1.36009 | 0.15744 |
| BABA 6h | 1181.01 | 166.895 | 1.50353 | 0.17869 |
| Chitin 6h | 1097.03 | 87.0704 | 1.49281 | 0.0983465 |
| Epi 6h | 1109.7 | 77.8996 | 1.42739 | 0.0859159 |
| SA 6h | 1033.42 | 59.7955 | 1.52237 | 0.176074 |
| Me-JA 6h | 1026.19 | 177.221 | 1.45106 | 0.16488 |
Source Transcript PGSC0003DMT400034372 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G38620.1 | +3 | 1e-101 | 305 | 172/286 (60%) | myb domain protein 4 | chr4:18053866-18054876 FORWARD LENGTH=282 |
