Probe CUST_4965_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_4965_PI426222305 | JHI_St_60k_v1 | DMT400009056 | ATTTTGCCACTGCTCATGCCTATCCTGATCAATGGTAAATTTTTCAGGACCACATTGCAC |
All Microarray Probes Designed to Gene DMG400003521
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_4965_PI426222305 | JHI_St_60k_v1 | DMT400009056 | ATTTTGCCACTGCTCATGCCTATCCTGATCAATGGTAAATTTTTCAGGACCACATTGCAC |
| CUST_5179_PI426222305 | JHI_St_60k_v1 | DMT400009055 | CCCTCTTAGAGATGATTATGGTTCATCCAGGTGGGTTAATAGAGATTATTACTTATAGTA |
Microarray Signals from CUST_4965_PI426222305
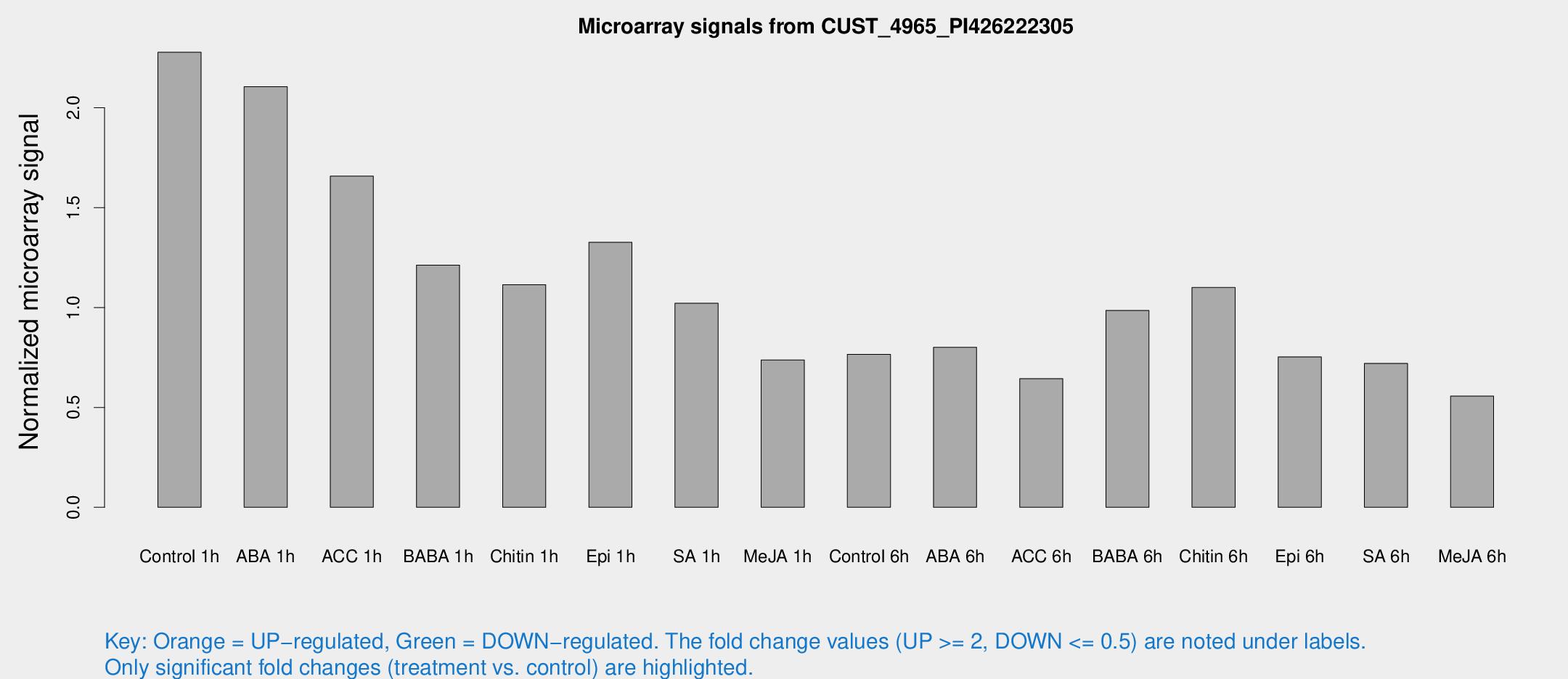
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 1539.33 | 250.938 | 2.27817 | 0.220346 |
| ABA 1h | 1320.38 | 364.273 | 2.10537 | 0.437676 |
| ACC 1h | 1272.16 | 433.067 | 1.6578 | 0.515816 |
| BABA 1h | 769.663 | 44.588 | 1.21271 | 0.131869 |
| Chitin 1h | 669.639 | 81.6753 | 1.11448 | 0.0645642 |
| Epi 1h | 762.644 | 70.9655 | 1.32671 | 0.108585 |
| SA 1h | 699.227 | 75.1049 | 1.0215 | 0.1766 |
| Me-JA 1h | 397.686 | 23.3683 | 0.73763 | 0.106997 |
| Control 6h | 653.506 | 277.761 | 0.76604 | 0.427973 |
| ABA 6h | 562.532 | 35.8936 | 0.80081 | 0.0866696 |
| ACC 6h | 516.954 | 127.287 | 0.644111 | 0.082395 |
| BABA 6h | 749.408 | 137.7 | 0.985374 | 0.161918 |
| Chitin 6h | 774.133 | 52.3192 | 1.10034 | 0.0791906 |
| Epi 6h | 570.744 | 85.9474 | 0.753218 | 0.121424 |
| SA 6h | 489.675 | 94.78 | 0.720287 | 0.0724096 |
| Me-JA 6h | 380.891 | 75.9543 | 0.556455 | 0.0722641 |
Source Transcript PGSC0003DMT400009056 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G66460.1 | +1 | 2e-144 | 418 | 197/285 (69%) | Glycosyl hydrolase superfamily protein | chr5:26538911-26540837 REVERSE LENGTH=431 |
