Probe CUST_49430_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_49430_PI426222305 | JHI_St_60k_v1 | DMT400051190 | AAAGACTTGCAAGAGCCTGTTCGTATTCCAGGATGCAGACCTATACACGGAAAGGATCTT |
All Microarray Probes Designed to Gene DMG400019882
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_49430_PI426222305 | JHI_St_60k_v1 | DMT400051190 | AAAGACTTGCAAGAGCCTGTTCGTATTCCAGGATGCAGACCTATACACGGAAAGGATCTT |
| CUST_49441_PI426222305 | JHI_St_60k_v1 | DMT400051191 | TTAAATAAATCTTTCTTTAACCGCAGGATGCAGACCTATACACGGAAAGGATCTTCCAGA |
Microarray Signals from CUST_49430_PI426222305
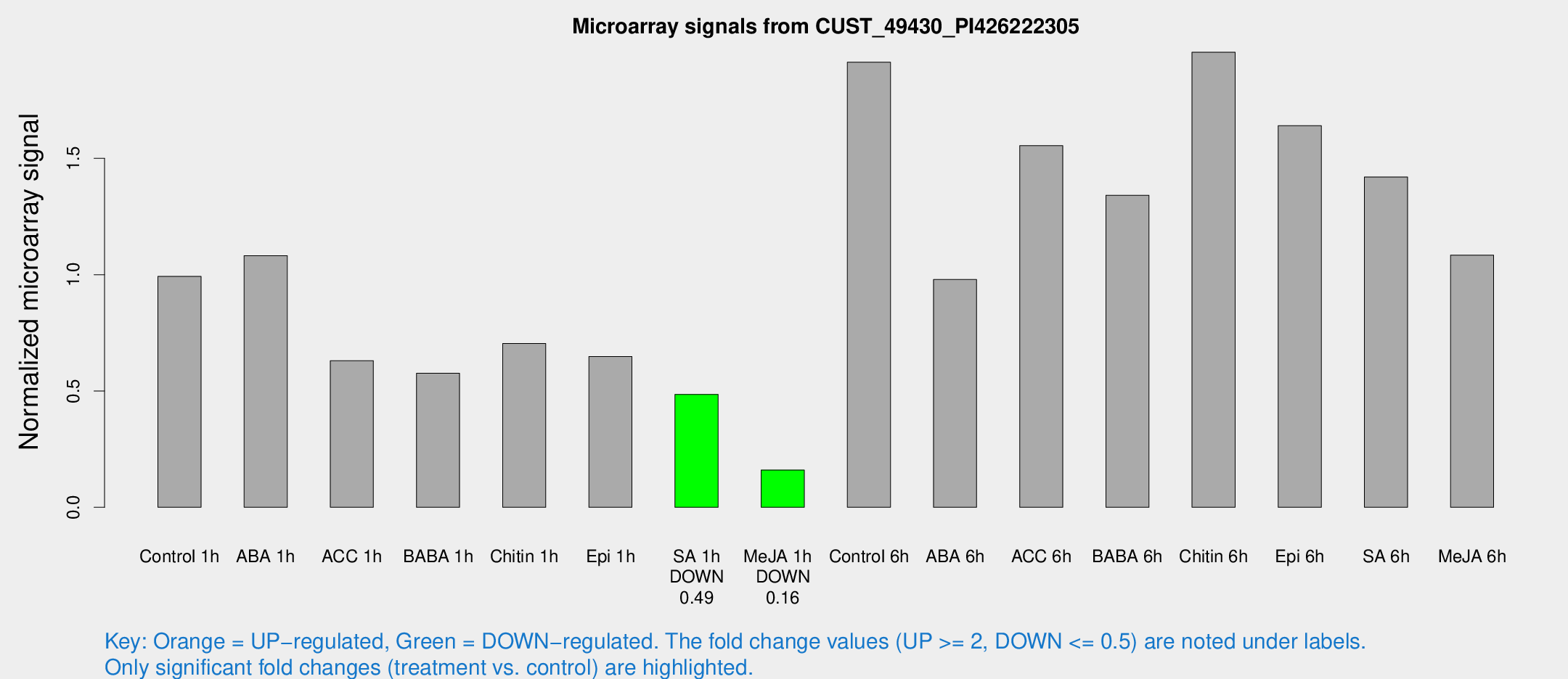
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 411.462 | 34.6218 | 0.992367 | 0.0577829 |
| ABA 1h | 393.985 | 22.9386 | 1.08196 | 0.0629807 |
| ACC 1h | 276.976 | 56.0076 | 0.629928 | 0.0868119 |
| BABA 1h | 248.554 | 65.6854 | 0.57632 | 0.119491 |
| Chitin 1h | 265.415 | 35.3303 | 0.704028 | 0.0420495 |
| Epi 1h | 233.705 | 24.8438 | 0.648404 | 0.0572182 |
| SA 1h | 207.427 | 21.9304 | 0.485434 | 0.0333744 |
| Me-JA 1h | 54.7642 | 7.27471 | 0.160125 | 0.0192893 |
| Control 6h | 817.524 | 142.362 | 1.91367 | 0.202887 |
| ABA 6h | 429.791 | 25.1018 | 0.979127 | 0.0570173 |
| ACC 6h | 741.27 | 72.2648 | 1.55477 | 0.105519 |
| BABA 6h | 650.988 | 155.595 | 1.3415 | 0.297962 |
| Chitin 6h | 857.658 | 49.6719 | 1.9563 | 0.113224 |
| Epi 6h | 774.023 | 95.6267 | 1.64073 | 0.318399 |
| SA 6h | 579.19 | 33.67 | 1.41979 | 0.203248 |
| Me-JA 6h | 516.723 | 197.924 | 1.08418 | 0.35754 |
Source Transcript PGSC0003DMT400051190 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G01070.1 | +3 | 0.0 | 523 | 268/469 (57%) | UDP-Glycosyltransferase superfamily protein | chr4:461858-463300 REVERSE LENGTH=480 |
