Probe CUST_4931_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_4931_PI426222305 | JHI_St_60k_v1 | DMT400009390 | GTGAATTCCATCGAGGCACTAGCAGAAGGAAGCTCAGCAAACTTAGGAGAAATCAACTTG |
All Microarray Probes Designed to Gene DMG400003649
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_4964_PI426222305 | JHI_St_60k_v1 | DMT400009388 | TAGCTGCAGGTGATGCGTGCAATAAATTAACTAGTTGATGTAACTTGTATGGTTGTTAAT |
| CUST_4931_PI426222305 | JHI_St_60k_v1 | DMT400009390 | GTGAATTCCATCGAGGCACTAGCAGAAGGAAGCTCAGCAAACTTAGGAGAAATCAACTTG |
Microarray Signals from CUST_4931_PI426222305
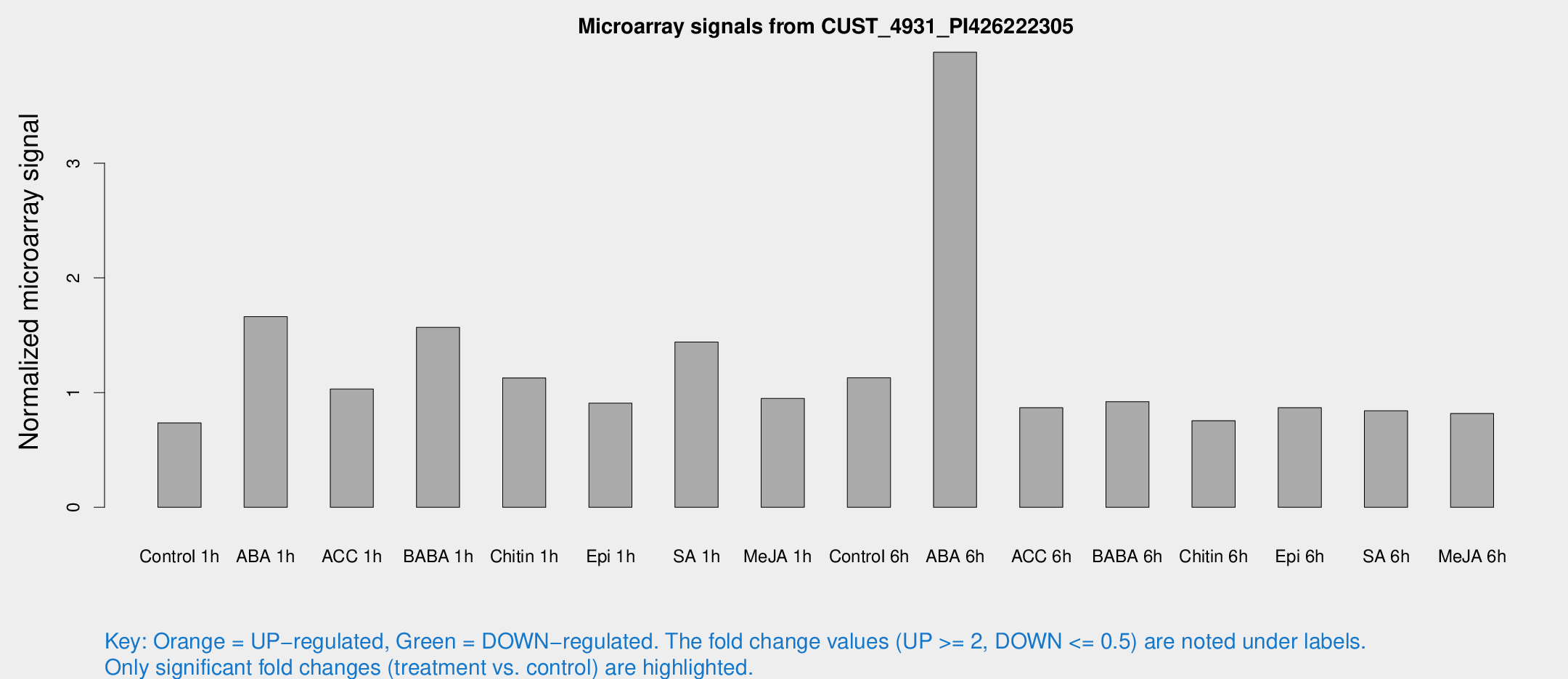
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 8.03968 | 3.60344 | 0.735542 | 0.364004 |
| ABA 1h | 16.8436 | 5.32593 | 1.66166 | 0.448499 |
| ACC 1h | 11.7304 | 4.04765 | 1.03108 | 0.413702 |
| BABA 1h | 16.1105 | 3.9989 | 1.56818 | 0.449 |
| Chitin 1h | 11.7583 | 4.04779 | 1.12803 | 0.490916 |
| Epi 1h | 8.33231 | 3.59312 | 0.907542 | 0.417062 |
| SA 1h | 17.21 | 5.53716 | 1.4404 | 0.616481 |
| Me-JA 1h | 8.31326 | 3.80983 | 0.948398 | 0.468435 |
| Control 6h | 12.668 | 3.99121 | 1.12918 | 0.434166 |
| ABA 6h | 47.6446 | 12.2739 | 3.96735 | 1.09997 |
| ACC 6h | 11.1807 | 4.66827 | 0.868093 | 0.413501 |
| BABA 6h | 12.1255 | 4.51422 | 0.921346 | 0.411221 |
| Chitin 6h | 8.39011 | 4.33848 | 0.754723 | 0.39759 |
| Epi 6h | 10.6537 | 4.47924 | 0.867941 | 0.391668 |
| SA 6h | 8.7264 | 4.12616 | 0.842062 | 0.407294 |
| Me-JA 6h | 8.71669 | 3.97497 | 0.817282 | 0.398693 |
Source Transcript PGSC0003DMT400009390 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G38580.1 | +1 | 9e-72 | 221 | 107/159 (67%) | farnesylated protein 6 | chr4:18034596-18035693 FORWARD LENGTH=153 |
