Probe CUST_49213_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_49213_PI426222305 | JHI_St_60k_v1 | DMT400044066 | TAAGCAGGAGATTAAGAAACCTGAGGTCAAGACCATTGAGATTAATTAGTGGTTAATTAC |
All Microarray Probes Designed to Gene DMG400017098
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_49213_PI426222305 | JHI_St_60k_v1 | DMT400044066 | TAAGCAGGAGATTAAGAAACCTGAGGTCAAGACCATTGAGATTAATTAGTGGTTAATTAC |
Microarray Signals from CUST_49213_PI426222305
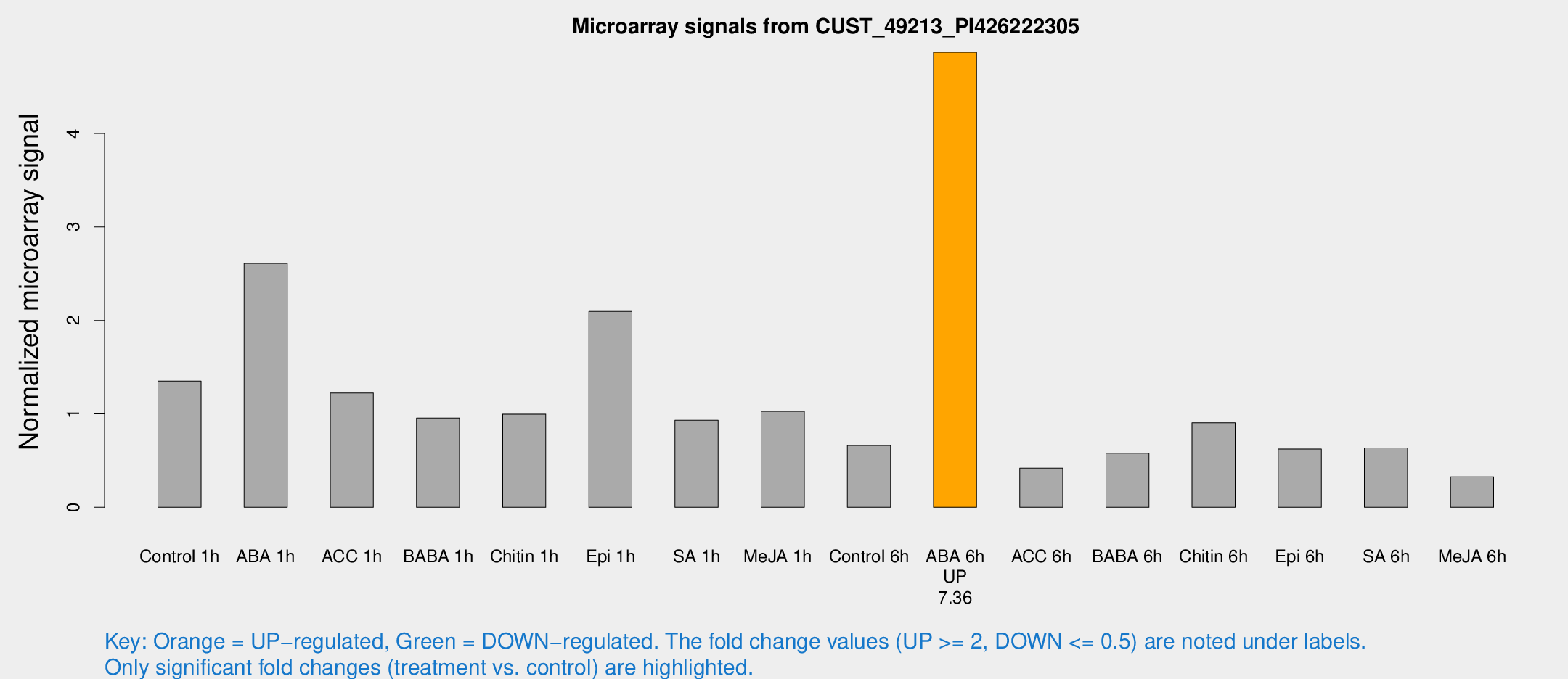
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 28.5652 | 3.85659 | 1.35113 | 0.182046 |
| ABA 1h | 53.0364 | 14.8972 | 2.60935 | 0.743239 |
| ACC 1h | 27.3191 | 4.94058 | 1.22263 | 0.243179 |
| BABA 1h | 19.4316 | 3.89084 | 0.955124 | 0.191005 |
| Chitin 1h | 19.7536 | 3.8632 | 0.996903 | 0.258766 |
| Epi 1h | 41.7978 | 12.7676 | 2.09517 | 0.598718 |
| SA 1h | 21.2246 | 4.50583 | 0.93145 | 0.212675 |
| Me-JA 1h | 20.8658 | 6.85326 | 1.0268 | 0.669844 |
| Control 6h | 15.4688 | 4.61275 | 0.661563 | 0.212382 |
| ABA 6h | 111.963 | 17.9855 | 4.86872 | 0.632847 |
| ACC 6h | 10.7165 | 4.25625 | 0.418869 | 0.180204 |
| BABA 6h | 13.7487 | 4.10576 | 0.578883 | 0.174928 |
| Chitin 6h | 20.8715 | 4.19661 | 0.905265 | 0.192423 |
| Epi 6h | 15.8034 | 4.35671 | 0.624114 | 0.217338 |
| SA 6h | 14.3319 | 4.03837 | 0.634325 | 0.283119 |
| Me-JA 6h | 6.93849 | 3.61479 | 0.326397 | 0.173711 |
Source Transcript PGSC0003DMT400044066 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G59720.1 | +2 | 6e-36 | 127 | 80/137 (58%) | heat shock protein 18.2 | chr5:24062632-24063117 FORWARD LENGTH=161 |
