Probe CUST_49016_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_49016_PI426222305 | JHI_St_60k_v1 | DMT400021857 | AGTTCAGGTAGTACTTCTAACACAAATAGCCCATTGATCCACAACAGAAGGGCAATTTGA |
All Microarray Probes Designed to Gene DMG402008483
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_49016_PI426222305 | JHI_St_60k_v1 | DMT400021857 | AGTTCAGGTAGTACTTCTAACACAAATAGCCCATTGATCCACAACAGAAGGGCAATTTGA |
Microarray Signals from CUST_49016_PI426222305
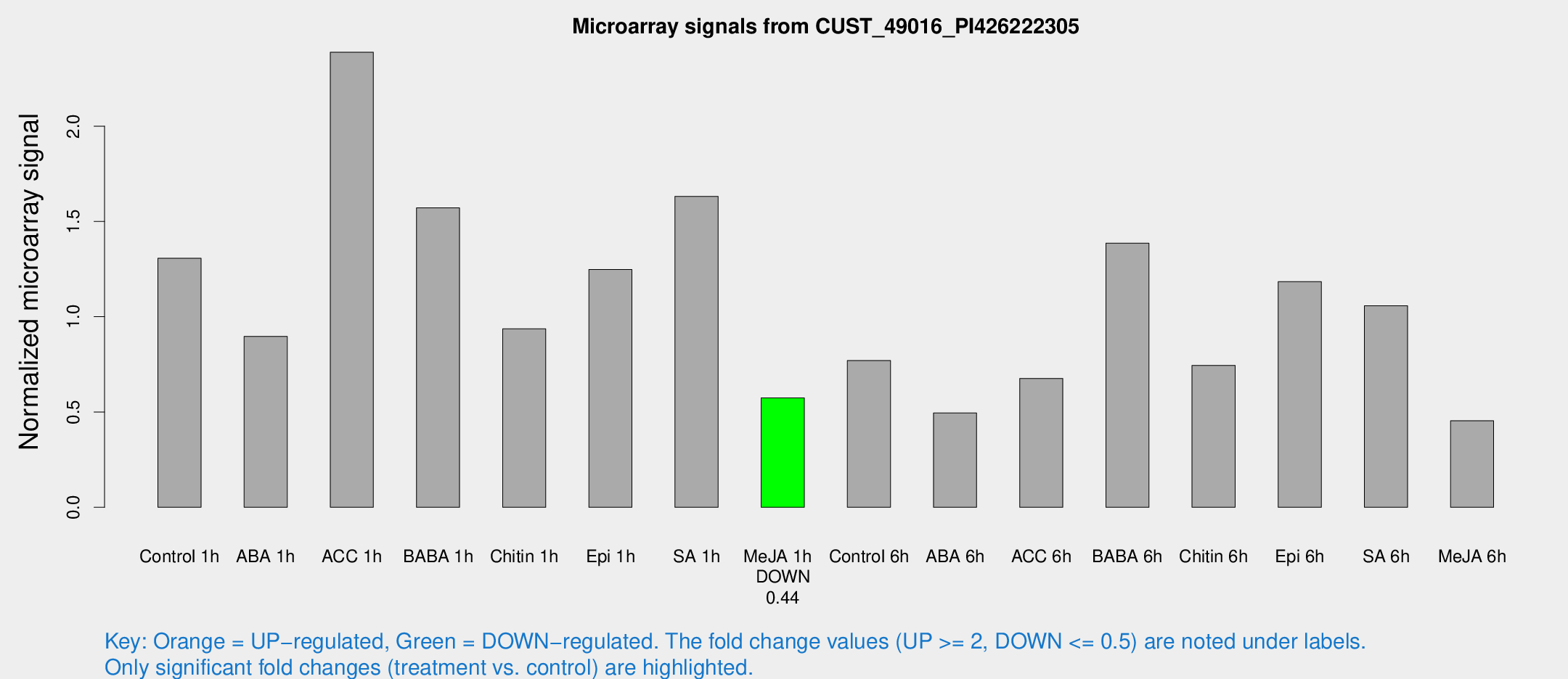
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 56.427 | 12.5907 | 1.30639 | 0.21928 |
| ABA 1h | 32.8151 | 3.59499 | 0.897014 | 0.099476 |
| ACC 1h | 113.078 | 35.1034 | 2.38781 | 0.715613 |
| BABA 1h | 62.3511 | 4.99894 | 1.57127 | 0.125964 |
| Chitin 1h | 35.9014 | 7.16234 | 0.936351 | 0.172421 |
| Epi 1h | 44.8982 | 5.17987 | 1.24784 | 0.1214 |
| SA 1h | 69.0918 | 5.14606 | 1.63118 | 0.120912 |
| Me-JA 1h | 19.4279 | 3.45963 | 0.574509 | 0.107208 |
| Control 6h | 40.223 | 19.4245 | 0.769697 | 0.381909 |
| ABA 6h | 21.7162 | 3.67712 | 0.494846 | 0.0849212 |
| ACC 6h | 33.2821 | 7.40628 | 0.676329 | 0.200651 |
| BABA 6h | 66.0922 | 13.2894 | 1.38591 | 0.248653 |
| Chitin 6h | 33.8886 | 7.05809 | 0.744884 | 0.129324 |
| Epi 6h | 72.7824 | 35.6588 | 1.18437 | 0.68466 |
| SA 6h | 45.7769 | 11.8302 | 1.05745 | 0.18486 |
| Me-JA 6h | 22.4232 | 7.76763 | 0.453738 | 0.321098 |
Source Transcript PGSC0003DMT400021857 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G21270.1 | +1 | 2e-169 | 503 | 264/540 (49%) | wall-associated kinase 2 | chr1:7444997-7447345 FORWARD LENGTH=732 |
