Probe CUST_48986_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_48986_PI426222305 | JHI_St_60k_v1 | DMT400056253 | ACGTACCTCTAGGTGAACTCATGGCTTGTCTTGAAATACAATATTATTACTTTCTCTGAT |
All Microarray Probes Designed to Gene DMG400021856
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_48986_PI426222305 | JHI_St_60k_v1 | DMT400056253 | ACGTACCTCTAGGTGAACTCATGGCTTGTCTTGAAATACAATATTATTACTTTCTCTGAT |
| CUST_48981_PI426222305 | JHI_St_60k_v1 | DMT400056252 | GCTCATATTAGTGGTGAATTGTGTCCTATTTTGAAAAATGCAGGGGCCTTGTGGCTATGA |
Microarray Signals from CUST_48986_PI426222305
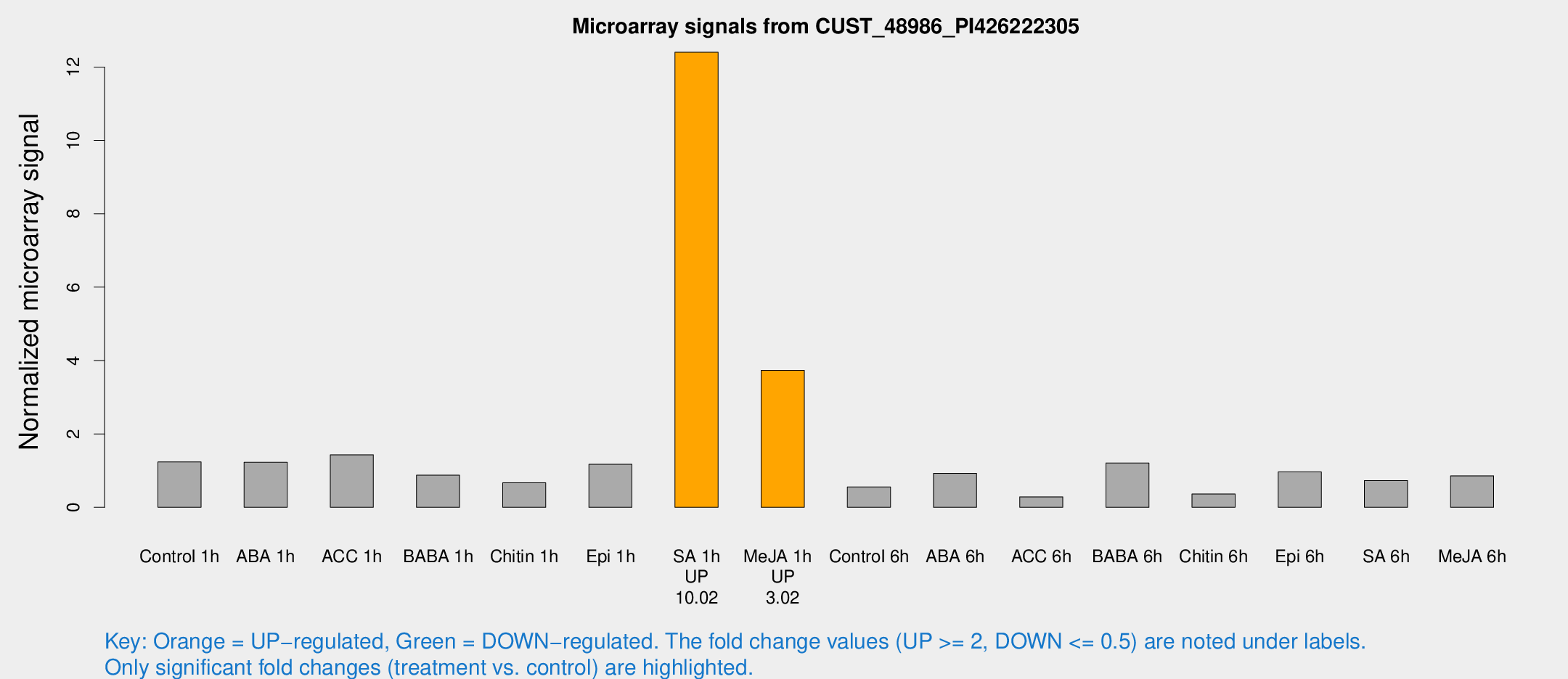
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 33.3614 | 10.1574 | 1.23827 | 0.394136 |
| ABA 1h | 26.5644 | 3.69581 | 1.22596 | 0.165976 |
| ACC 1h | 39.1904 | 10.8951 | 1.43074 | 0.410838 |
| BABA 1h | 21.0296 | 4.32163 | 0.874632 | 0.175254 |
| Chitin 1h | 14.8233 | 3.25571 | 0.668478 | 0.152514 |
| Epi 1h | 24.5896 | 3.40628 | 1.17215 | 0.167891 |
| SA 1h | 315.823 | 54.0148 | 12.4035 | 1.83212 |
| Me-JA 1h | 78.6065 | 21.9582 | 3.73351 | 0.695011 |
| Control 6h | 15.6097 | 5.03883 | 0.555714 | 0.204124 |
| ABA 6h | 28.3388 | 10.561 | 0.924749 | 0.4796 |
| ACC 6h | 8.02058 | 4.01934 | 0.284938 | 0.143827 |
| BABA 6h | 47.5457 | 20.0379 | 1.20855 | 1.43834 |
| Chitin 6h | 11.1957 | 5.07793 | 0.363566 | 0.168371 |
| Epi 6h | 26.5619 | 4.11295 | 0.966676 | 0.154664 |
| SA 6h | 19.4361 | 5.64306 | 0.729662 | 0.36475 |
| Me-JA 6h | 21.2669 | 3.95582 | 0.857799 | 0.223984 |
Source Transcript PGSC0003DMT400056253 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G28480.1 | +2 | 3e-36 | 127 | 63/106 (59%) | Thioredoxin superfamily protein | chr1:10013634-10014047 REVERSE LENGTH=137 |
