Probe CUST_48982_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_48982_PI426222305 | JHI_St_60k_v1 | DMT400056272 | ATAGCAATGGTTGATCCTAGTGGTGTCCTTAAGAAGTCCGGGATTGCTGAATGCCTTCAA |
All Microarray Probes Designed to Gene DMG400021866
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_48982_PI426222305 | JHI_St_60k_v1 | DMT400056272 | ATAGCAATGGTTGATCCTAGTGGTGTCCTTAAGAAGTCCGGGATTGCTGAATGCCTTCAA |
| CUST_48976_PI426222305 | JHI_St_60k_v1 | DMT400056271 | GGAAATCCTTGTGGAAATGTTTTTACGTCTGAACACTTGCAAGAGATTGCTGAATGCCTT |
Microarray Signals from CUST_48982_PI426222305
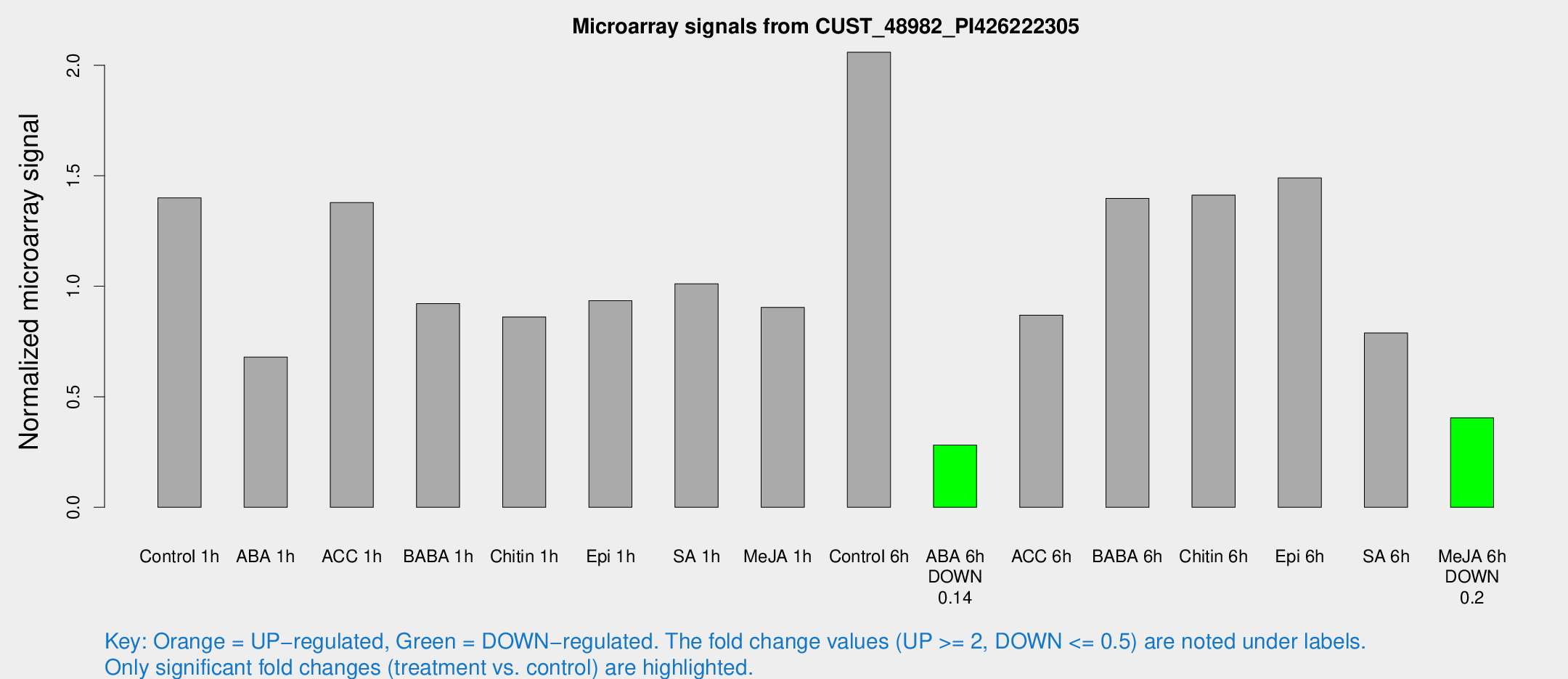
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 251.243 | 33.4253 | 1.3996 | 0.104481 |
| ABA 1h | 120.643 | 36.7228 | 0.679317 | 0.309764 |
| ACC 1h | 255.182 | 40.0956 | 1.37847 | 0.162086 |
| BABA 1h | 157.868 | 14.2831 | 0.921808 | 0.0578541 |
| Chitin 1h | 144.328 | 34.4343 | 0.861205 | 0.255132 |
| Epi 1h | 144.816 | 17.8451 | 0.935159 | 0.123959 |
| SA 1h | 186.509 | 24.1317 | 1.01142 | 0.204618 |
| Me-JA 1h | 132.551 | 18.7767 | 0.904469 | 0.152187 |
| Control 6h | 416.527 | 142.91 | 2.05876 | 0.675526 |
| ABA 6h | 54.8021 | 9.82007 | 0.281654 | 0.0751366 |
| ACC 6h | 211.442 | 94.0604 | 0.869372 | 0.259298 |
| BABA 6h | 301.373 | 94.0985 | 1.39779 | 0.434447 |
| Chitin 6h | 270.181 | 39.2065 | 1.41253 | 0.154603 |
| Epi 6h | 320.219 | 79.3989 | 1.49013 | 0.585285 |
| SA 6h | 151.606 | 41.0344 | 0.78849 | 0.176714 |
| Me-JA 6h | 72.5652 | 10.4401 | 0.405059 | 0.0327214 |
Source Transcript PGSC0003DMT400056272 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G53970.1 | +1 | 5e-165 | 477 | 232/402 (58%) | Tyrosine transaminase family protein | chr5:21910676-21912594 FORWARD LENGTH=414 |
