Probe CUST_48598_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_48598_PI426222305 | JHI_St_60k_v1 | DMT400035991 | AGACCCGCTTTTCCTCGTGGGATCATCAACCATGTTTACTAAGTTGTCAAATGCGGTTGT |
All Microarray Probes Designed to Gene DMG400013857
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_48592_PI426222305 | JHI_St_60k_v1 | DMT400035989 | TTTAAGGTGTTGAGAGAGGGAAATAAAGGTTACGGAATCTTAGTTTCTTCTGCTCCAAAA |
| CUST_48598_PI426222305 | JHI_St_60k_v1 | DMT400035991 | AGACCCGCTTTTCCTCGTGGGATCATCAACCATGTTTACTAAGTTGTCAAATGCGGTTGT |
Microarray Signals from CUST_48598_PI426222305
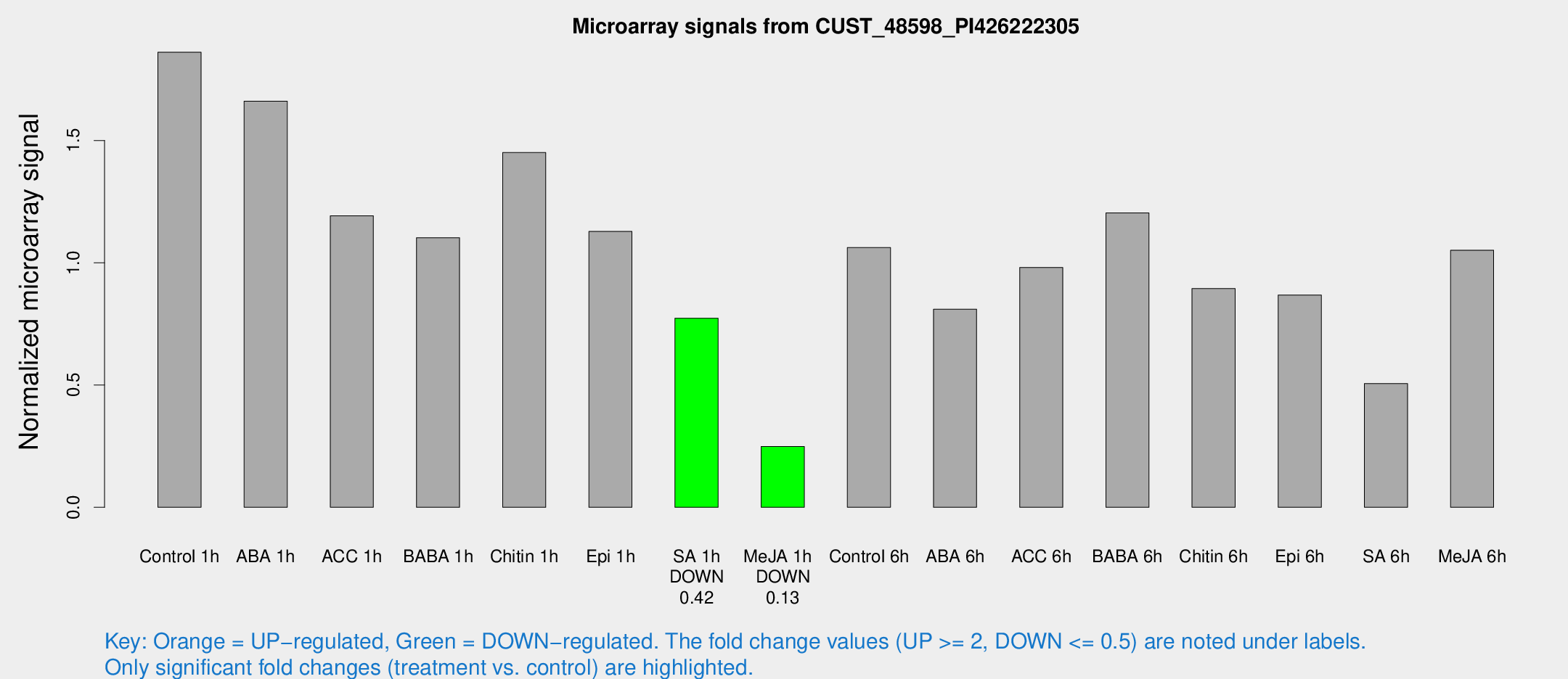
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 79.6691 | 10.7953 | 1.861 | 0.139363 |
| ABA 1h | 62.7982 | 8.60744 | 1.66101 | 0.12739 |
| ACC 1h | 52.6938 | 7.82541 | 1.19222 | 0.267889 |
| BABA 1h | 50.1601 | 16.3496 | 1.10233 | 0.299782 |
| Chitin 1h | 56.9619 | 10.8002 | 1.45082 | 0.242914 |
| Epi 1h | 41.3788 | 4.00937 | 1.12794 | 0.111575 |
| SA 1h | 35.6639 | 8.32841 | 0.773163 | 0.181511 |
| Me-JA 1h | 9.07715 | 3.04886 | 0.248944 | 0.0952728 |
| Control 6h | 52.8947 | 17.9748 | 1.06227 | 0.418202 |
| ABA 6h | 37.3692 | 6.40061 | 0.81013 | 0.10906 |
| ACC 6h | 48.0306 | 6.04194 | 0.98071 | 0.100601 |
| BABA 6h | 58.6282 | 11.5049 | 1.20355 | 0.215378 |
| Chitin 6h | 40.1493 | 4.07492 | 0.894345 | 0.0912284 |
| Epi 6h | 43.3842 | 10.5823 | 0.867989 | 0.204174 |
| SA 6h | 28.7567 | 11.5343 | 0.505711 | 0.335923 |
| Me-JA 6h | 45.2156 | 7.21086 | 1.05135 | 0.125239 |
Source Transcript PGSC0003DMT400035991 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G51460.3 | +3 | 4e-174 | 509 | 268/413 (65%) | Haloacid dehalogenase-like hydrolase (HAD) superfamily protein | chr5:20902266-20904292 FORWARD LENGTH=385 |
