Probe CUST_48523_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_48523_PI426222305 | JHI_St_60k_v1 | DMT400043381 | GGAAGCATCCTAAGTCATGTAAATGTTTGTCTTTTGATTGTCACAATCTGAACAAACATC |
All Microarray Probes Designed to Gene DMG400016842
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_48526_PI426222305 | JHI_St_60k_v1 | DMT400043382 | GGAAGCATCCTAAGTCATGTAAATGTTTGTCTTTTGATTGTCACAATCTGAACAAACATC |
| CUST_48523_PI426222305 | JHI_St_60k_v1 | DMT400043381 | GGAAGCATCCTAAGTCATGTAAATGTTTGTCTTTTGATTGTCACAATCTGAACAAACATC |
Microarray Signals from CUST_48523_PI426222305
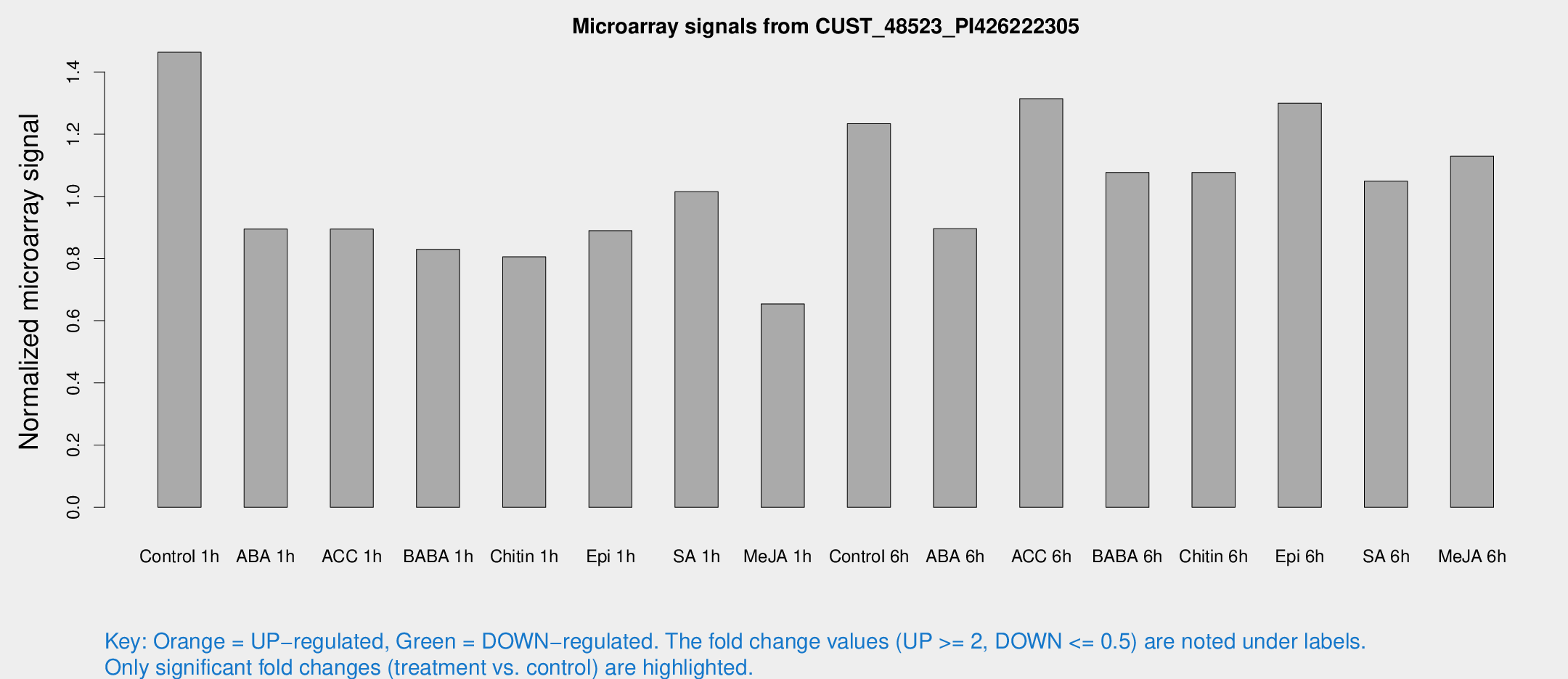
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 251.615 | 59.5764 | 1.46342 | 0.238717 |
| ABA 1h | 129.328 | 8.22729 | 0.894626 | 0.0648025 |
| ACC 1h | 157.278 | 31.6365 | 0.89479 | 0.138226 |
| BABA 1h | 136.545 | 26.3796 | 0.829549 | 0.0959375 |
| Chitin 1h | 118.551 | 7.76144 | 0.805482 | 0.0525584 |
| Epi 1h | 129.009 | 19.5399 | 0.889607 | 0.132977 |
| SA 1h | 175.389 | 31.4326 | 1.01497 | 0.132301 |
| Me-JA 1h | 87.8721 | 7.47534 | 0.654165 | 0.0468839 |
| Control 6h | 207.174 | 31.7736 | 1.23381 | 0.106203 |
| ABA 6h | 155.918 | 9.83633 | 0.895965 | 0.0564122 |
| ACC 6h | 250.995 | 34.3291 | 1.31398 | 0.0936316 |
| BABA 6h | 198.995 | 20.8486 | 1.07689 | 0.0712425 |
| Chitin 6h | 188.485 | 16.2699 | 1.07682 | 0.121283 |
| Epi 6h | 246.046 | 39.4347 | 1.2997 | 0.162602 |
| SA 6h | 176.957 | 36.9584 | 1.049 | 0.131371 |
| Me-JA 6h | 189.035 | 30.306 | 1.12924 | 0.108035 |
Source Transcript PGSC0003DMT400043381 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G32910.1 | +3 | 9e-56 | 207 | 152/418 (36%) | DCD (Development and Cell Death) domain protein | chr2:13959685-13961989 FORWARD LENGTH=691 |
