Probe CUST_47837_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_47837_PI426222305 | JHI_St_60k_v1 | DMT400068327 | TCGTTAAAGCTAGTGGTGATGTTGATGCTCTCTACGGCATTCTTCTTTGCAACTGTCTAG |
All Microarray Probes Designed to Gene DMG400026565
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_47852_PI426222305 | JHI_St_60k_v1 | DMT400068328 | GGTGCCATGATTTTCATTCAGCAGTTTGTATCAAATGTTTTTCTACTGTGGTAAAGTCAT |
| CUST_47837_PI426222305 | JHI_St_60k_v1 | DMT400068327 | TCGTTAAAGCTAGTGGTGATGTTGATGCTCTCTACGGCATTCTTCTTTGCAACTGTCTAG |
Microarray Signals from CUST_47837_PI426222305
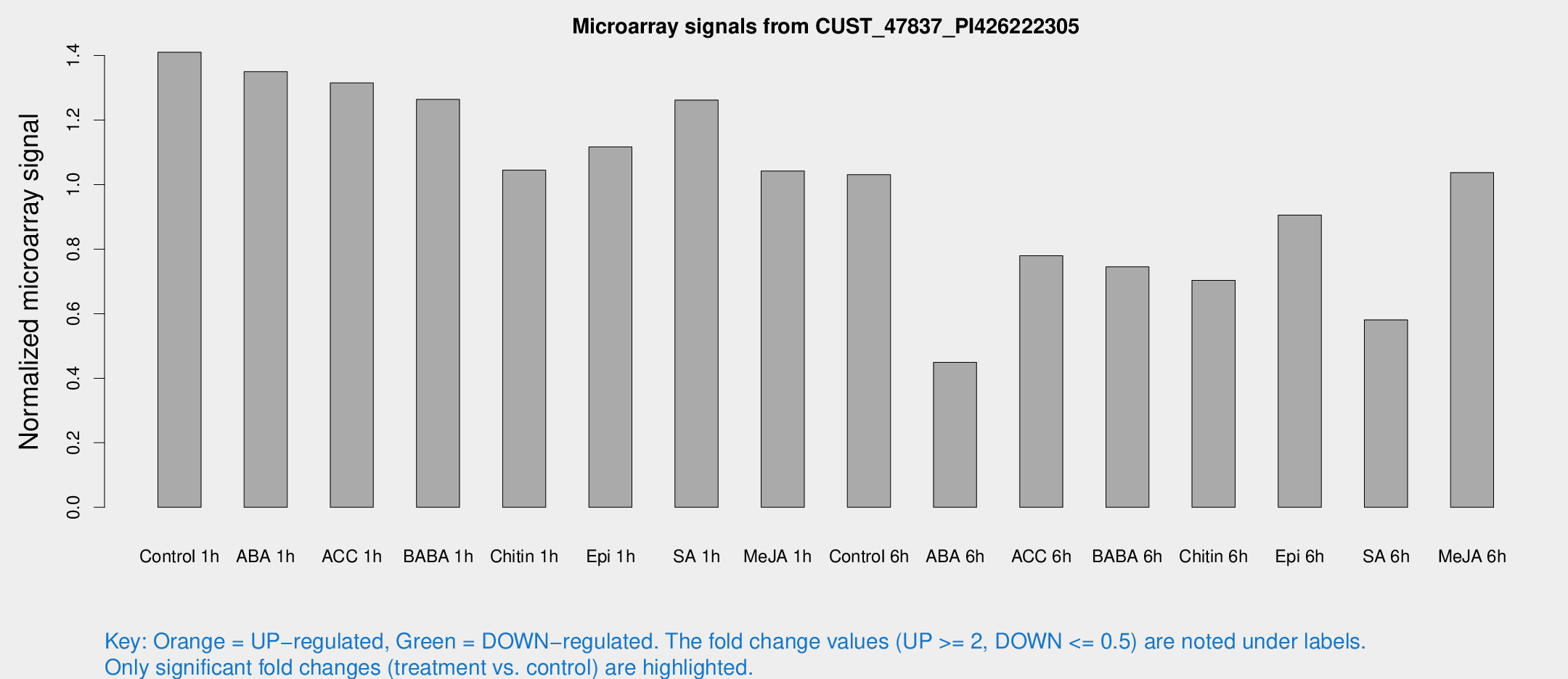
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 7324.97 | 1264.25 | 1.41023 | 0.152386 |
| ABA 1h | 6038.35 | 348.859 | 1.3499 | 0.0993784 |
| ACC 1h | 7254.21 | 1609.41 | 1.31533 | 0.241024 |
| BABA 1h | 6180.26 | 384.956 | 1.2639 | 0.151026 |
| Chitin 1h | 4819.6 | 597.219 | 1.04485 | 0.124851 |
| Epi 1h | 4907.33 | 318.504 | 1.11701 | 0.0644943 |
| SA 1h | 6680.81 | 954.611 | 1.26196 | 0.167994 |
| Me-JA 1h | 4301 | 248.413 | 1.04218 | 0.107866 |
| Control 6h | 5710.9 | 1526.55 | 1.03069 | 0.245127 |
| ABA 6h | 2450.36 | 308.923 | 0.449309 | 0.076489 |
| ACC 6h | 4672.65 | 891.519 | 0.779428 | 0.0450045 |
| BABA 6h | 4244.69 | 362.955 | 0.745143 | 0.0451836 |
| Chitin 6h | 3833.82 | 431.367 | 0.703374 | 0.0713408 |
| Epi 6h | 5255.5 | 728.205 | 0.905651 | 0.150488 |
| SA 6h | 3314.07 | 1001.34 | 0.580757 | 0.169432 |
| Me-JA 6h | 5227.61 | 302.278 | 1.03729 | 0.0695076 |
Source Transcript PGSC0003DMT400068327 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G60920.1 | +1 | 0.0 | 676 | 300/373 (80%) | COBRA-like extracellular glycosyl-phosphatidyl inositol-anchored protein family | chr5:24511466-24513932 REVERSE LENGTH=456 |
