Probe CUST_47466_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_47466_PI426222305 | JHI_St_60k_v1 | DMT400064777 | GGCACTAGTTCATCGTTAATTTTGACAATTGTGAGTGCAGATTGGCTTGAAGCACCAGCA |
All Microarray Probes Designed to Gene DMG400025160
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_47445_PI426222305 | JHI_St_60k_v1 | DMT400064776 | GTTTCCCTTCATTACTCTTTGGTTTGTTCAGGGGAGATTTTGGAAAATGGCTATCTTTGA |
| CUST_47466_PI426222305 | JHI_St_60k_v1 | DMT400064777 | GGCACTAGTTCATCGTTAATTTTGACAATTGTGAGTGCAGATTGGCTTGAAGCACCAGCA |
Microarray Signals from CUST_47466_PI426222305
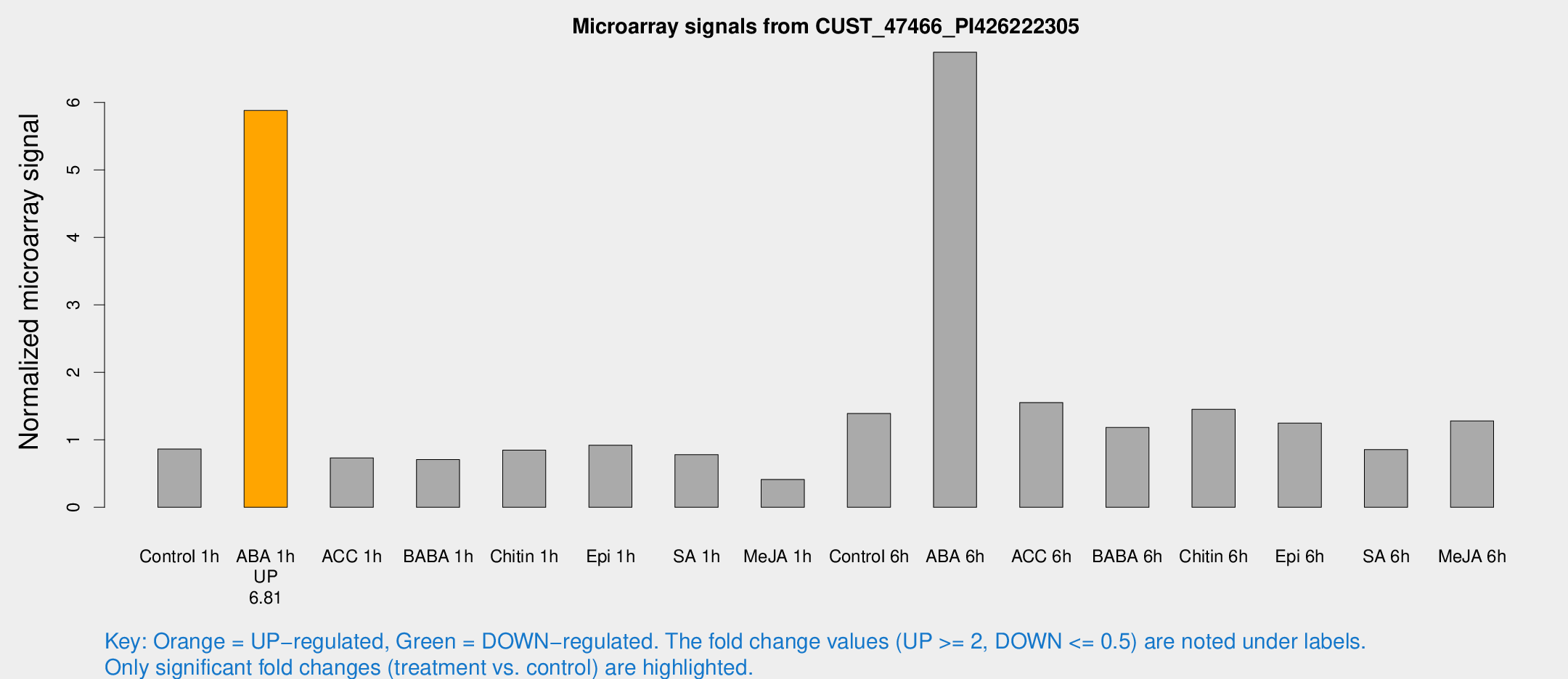
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 33.4678 | 3.70045 | 0.863473 | 0.0959041 |
| ABA 1h | 204.861 | 26.5861 | 5.88132 | 0.413032 |
| ACC 1h | 29.3651 | 3.85963 | 0.733022 | 0.0987218 |
| BABA 1h | 27.2331 | 4.80493 | 0.707296 | 0.104729 |
| Chitin 1h | 30.494 | 5.61383 | 0.847669 | 0.107305 |
| Epi 1h | 31.5583 | 4.56988 | 0.920905 | 0.13641 |
| SA 1h | 31.0015 | 3.66807 | 0.77924 | 0.0923357 |
| Me-JA 1h | 15.769 | 5.95791 | 0.412617 | 0.196393 |
| Control 6h | 57.8766 | 14.1385 | 1.38917 | 0.278276 |
| ABA 6h | 290.127 | 57.4099 | 6.74399 | 1.15947 |
| ACC 6h | 69.6316 | 7.21029 | 1.55208 | 0.126581 |
| BABA 6h | 53.0825 | 9.2127 | 1.18332 | 0.189722 |
| Chitin 6h | 59.9373 | 5.0683 | 1.45291 | 0.123037 |
| Epi 6h | 56.4002 | 10.2387 | 1.24817 | 0.314828 |
| SA 6h | 36.5405 | 10.967 | 0.855225 | 0.210179 |
| Me-JA 6h | 49.5858 | 4.45574 | 1.28107 | 0.115352 |
Source Transcript PGSC0003DMT400064777 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G50830.1 | +1 | 6e-50 | 170 | 99/192 (52%) | cold-regulated 413-plasma membrane 2 | chr3:18894109-18895355 REVERSE LENGTH=203 |
