Probe CUST_4715_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_4715_PI426222305 | JHI_St_60k_v1 | DMT400073853 | GGACTGATGAACCTGAAATCATGTATTGATTTTGAGTCCTTACTTGTCTAGTAACTCTTT |
All Microarray Probes Designed to Gene DMG400028700
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_4715_PI426222305 | JHI_St_60k_v1 | DMT400073853 | GGACTGATGAACCTGAAATCATGTATTGATTTTGAGTCCTTACTTGTCTAGTAACTCTTT |
Microarray Signals from CUST_4715_PI426222305
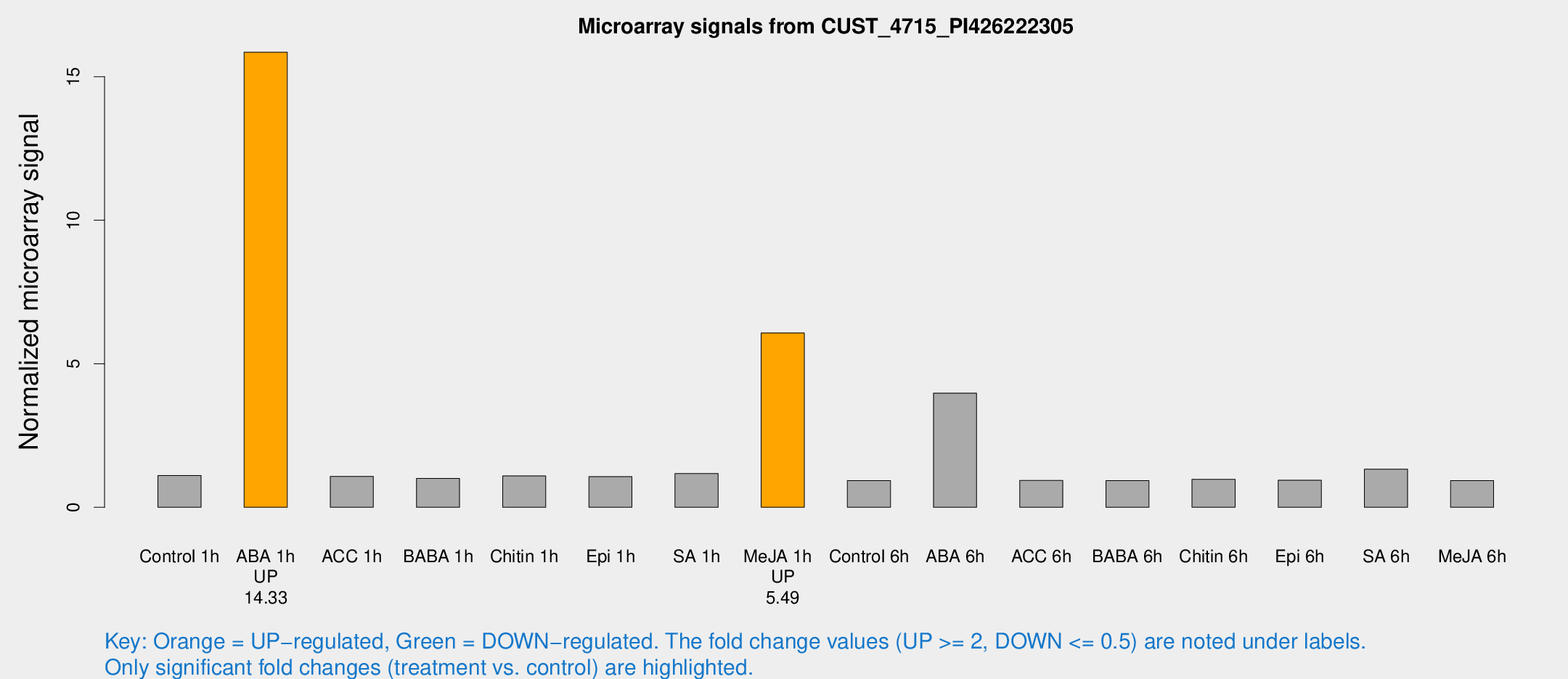
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 6.38718 | 3.00347 | 1.10619 | 0.549572 |
| ABA 1h | 86.6774 | 25.261 | 15.8524 | 4.10366 |
| ACC 1h | 6.24291 | 3.42178 | 1.07878 | 0.58567 |
| BABA 1h | 5.4508 | 3.16644 | 1.00406 | 0.582364 |
| Chitin 1h | 5.36075 | 2.9797 | 1.09374 | 0.58406 |
| Epi 1h | 5.06832 | 2.94328 | 1.067 | 0.602721 |
| SA 1h | 7.34078 | 3.01485 | 1.17582 | 0.564914 |
| Me-JA 1h | 28.7951 | 4.78752 | 6.07454 | 1.32468 |
| Control 6h | 5.26209 | 3.05394 | 0.933087 | 0.541173 |
| ABA 6h | 27.4471 | 8.53282 | 3.97832 | 1.62147 |
| ACC 6h | 6.17246 | 3.66102 | 0.935283 | 0.541595 |
| BABA 6h | 5.85914 | 3.39872 | 0.929235 | 0.538102 |
| Chitin 6h | 5.82059 | 3.37177 | 0.971935 | 0.562774 |
| Epi 6h | 6.04052 | 3.53217 | 0.943671 | 0.546439 |
| SA 6h | 7.98628 | 3.25936 | 1.32994 | 0.637012 |
| Me-JA 6h | 5.20637 | 3.01551 | 0.929511 | 0.538242 |
Source Transcript PGSC0003DMT400073853 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G18450.1 | +2 | 7e-36 | 131 | 69/89 (78%) | Integrase-type DNA-binding superfamily protein | chr5:6116097-6117020 REVERSE LENGTH=307 |
