Probe CUST_47147_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_47147_PI426222305 | JHI_St_60k_v1 | DMT400030417 | ATTGCTATCTTGGGATGCACTTTTTGTGTTGGTTTCCAGTTCTCACTCCGTCTCTATAAT |
All Microarray Probes Designed to Gene DMG401011647
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_47117_PI426222305 | JHI_St_60k_v1 | DMT400030415 | GATATGAAAAACAGCTTGCAACCAGGAAGAAATCATACATATCCTTTGGTGAAGGATTAG |
| CUST_47147_PI426222305 | JHI_St_60k_v1 | DMT400030417 | ATTGCTATCTTGGGATGCACTTTTTGTGTTGGTTTCCAGTTCTCACTCCGTCTCTATAAT |
Microarray Signals from CUST_47147_PI426222305
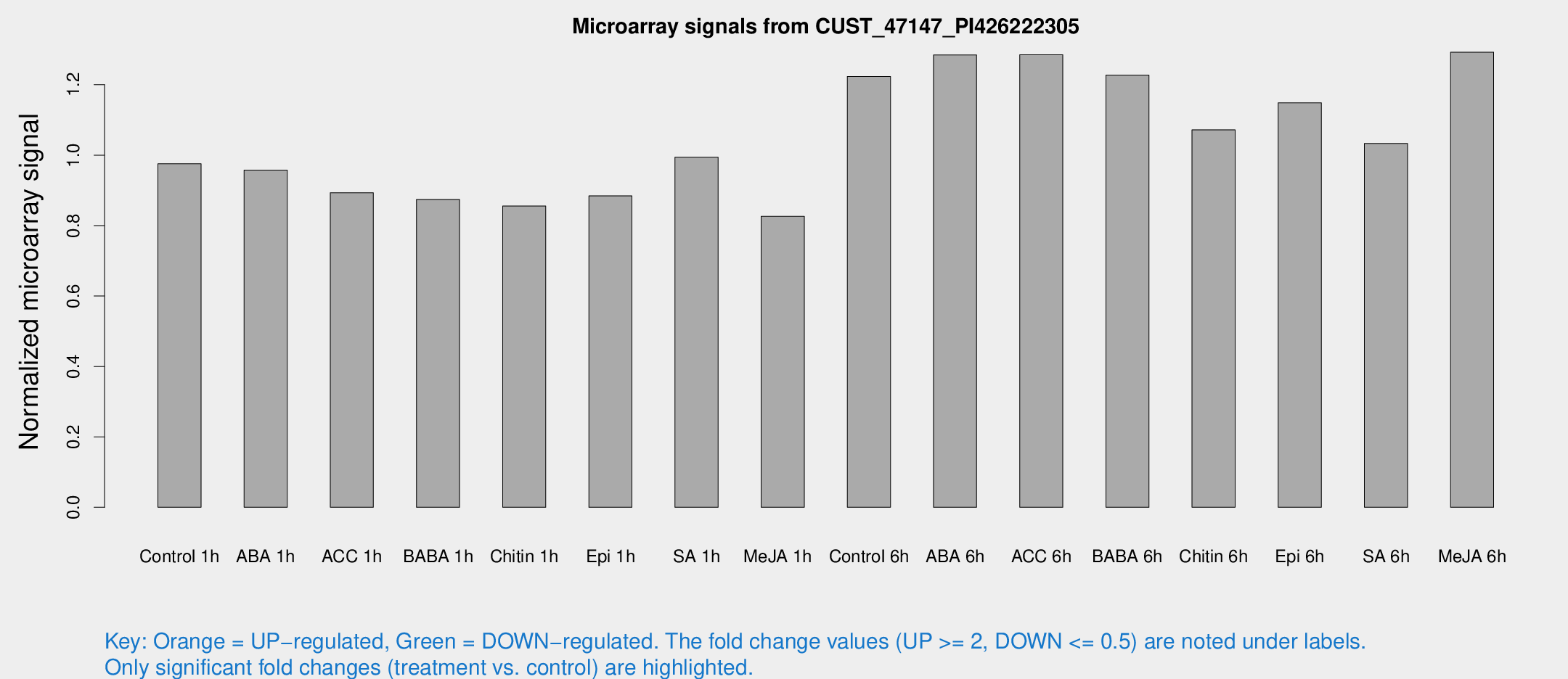
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 867.732 | 127.341 | 0.975573 | 0.0765871 |
| ABA 1h | 741.53 | 49.1858 | 0.957604 | 0.0554328 |
| ACC 1h | 836.923 | 162.639 | 0.893381 | 0.130687 |
| BABA 1h | 741.051 | 69.7285 | 0.874342 | 0.0506539 |
| Chitin 1h | 675.833 | 60.2958 | 0.855468 | 0.0701491 |
| Epi 1h | 671.017 | 51.0641 | 0.884258 | 0.051253 |
| SA 1h | 900.411 | 93.0327 | 0.994326 | 0.0730879 |
| Me-JA 1h | 588.813 | 34.2188 | 0.826125 | 0.0735813 |
| Control 6h | 1125.51 | 231.817 | 1.22346 | 0.180937 |
| ABA 6h | 1193.64 | 69.1961 | 1.28453 | 0.0742595 |
| ACC 6h | 1292.09 | 98.5254 | 1.28519 | 0.0848595 |
| BABA 6h | 1198.52 | 69.359 | 1.22743 | 0.0781698 |
| Chitin 6h | 1001.18 | 81.2495 | 1.07189 | 0.0620217 |
| Epi 6h | 1135.84 | 90.7118 | 1.14858 | 0.0863165 |
| SA 6h | 929.263 | 183.569 | 1.03315 | 0.114432 |
| Me-JA 6h | 1141.46 | 155.916 | 1.29226 | 0.0835876 |
Source Transcript PGSC0003DMT400030417 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G47030.1 | +2 | 6e-97 | 289 | 147/205 (72%) | ATPase, F1 complex, delta/epsilon subunit | chr5:19090384-19092034 FORWARD LENGTH=203 |
