Probe CUST_47116_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_47116_PI426222305 | JHI_St_60k_v1 | DMT400030402 | GTGTGTGTGTTTTAAGTGTGGAAAATGCCTATTTTGAGTTTGAGACGTGTGATTTTGTTT |
All Microarray Probes Designed to Gene DMG400011640
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_47116_PI426222305 | JHI_St_60k_v1 | DMT400030402 | GTGTGTGTGTTTTAAGTGTGGAAAATGCCTATTTTGAGTTTGAGACGTGTGATTTTGTTT |
Microarray Signals from CUST_47116_PI426222305
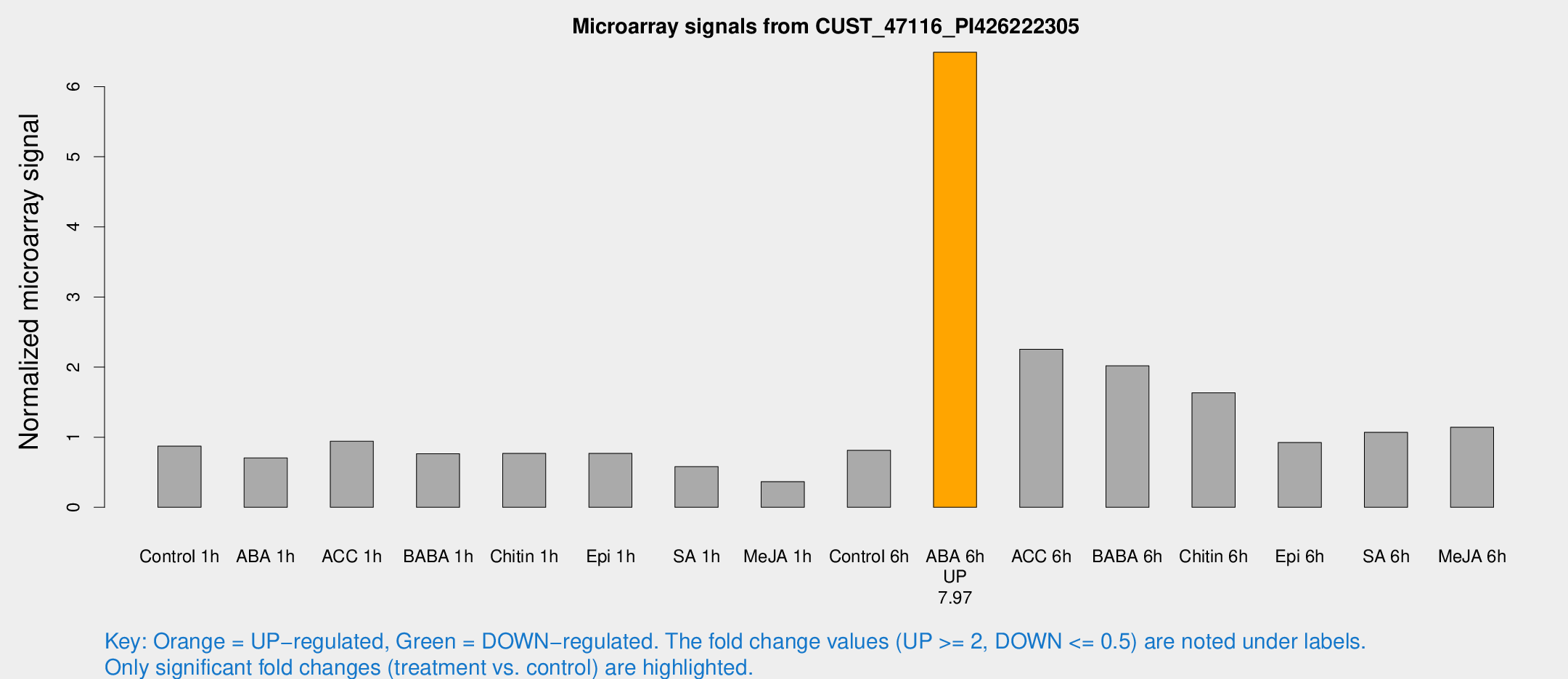
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 585.609 | 106.842 | 0.872563 | 0.108143 |
| ABA 1h | 433.481 | 104.068 | 0.703465 | 0.172749 |
| ACC 1h | 731.133 | 261.593 | 0.942398 | 0.341967 |
| BABA 1h | 480.809 | 38.771 | 0.76473 | 0.0544965 |
| Chitin 1h | 474.088 | 116.883 | 0.76811 | 0.124361 |
| Epi 1h | 451.011 | 88.2026 | 0.768919 | 0.165057 |
| SA 1h | 393.199 | 50.7726 | 0.581329 | 0.0500893 |
| Me-JA 1h | 216.049 | 64.1925 | 0.365677 | 0.116055 |
| Control 6h | 691.474 | 302.181 | 0.813996 | 0.469656 |
| ABA 6h | 4521.6 | 451.294 | 6.49126 | 0.408433 |
| ACC 6h | 1771.24 | 423.941 | 2.25488 | 0.29238 |
| BABA 6h | 1618.53 | 531.415 | 2.01886 | 0.583896 |
| Chitin 6h | 1148.77 | 162.117 | 1.63421 | 0.236613 |
| Epi 6h | 692.264 | 104.058 | 0.923071 | 0.223953 |
| SA 6h | 809.873 | 270.317 | 1.06976 | 0.38152 |
| Me-JA 6h | 738.763 | 42.8684 | 1.14245 | 0.14207 |
Source Transcript PGSC0003DMT400030402 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G05340.1 | +1 | 1e-166 | 474 | 234/298 (79%) | Peroxidase superfamily protein | chr5:1579142-1580819 REVERSE LENGTH=324 |
