Probe CUST_46777_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_46777_PI426222305 | JHI_St_60k_v1 | DMT400038285 | GCAGATAAAAATGGAGGCAACTGATAGTTAATTAATTCCCCTACATATTGAAGGTGCAAT |
All Microarray Probes Designed to Gene DMG400014773
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_46777_PI426222305 | JHI_St_60k_v1 | DMT400038285 | GCAGATAAAAATGGAGGCAACTGATAGTTAATTAATTCCCCTACATATTGAAGGTGCAAT |
Microarray Signals from CUST_46777_PI426222305
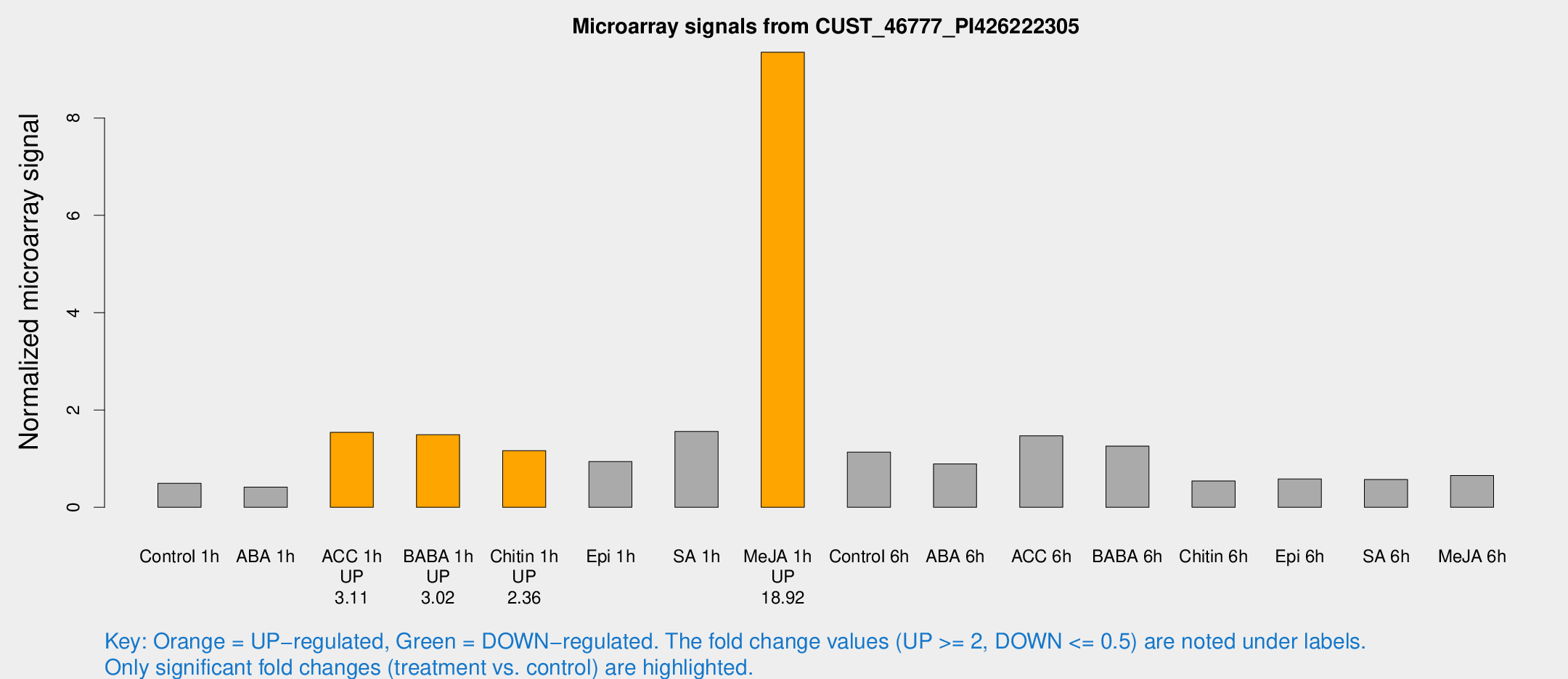
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 16.0806 | 3.95931 | 0.494255 | 0.111883 |
| ABA 1h | 14.4444 | 7.63192 | 0.414485 | 0.296939 |
| ACC 1h | 48.0651 | 4.21986 | 1.53931 | 0.217925 |
| BABA 1h | 45.5594 | 8.91464 | 1.49192 | 0.176407 |
| Chitin 1h | 31.8496 | 3.49014 | 1.16573 | 0.138304 |
| Epi 1h | 24.8821 | 3.27868 | 0.94088 | 0.125305 |
| SA 1h | 55.4559 | 20.9259 | 1.55779 | 0.589833 |
| Me-JA 1h | 238.277 | 39.8685 | 9.35189 | 0.838768 |
| Control 6h | 40.9485 | 13.5909 | 1.13513 | 0.457113 |
| ABA 6h | 31.8032 | 10.5483 | 0.8928 | 0.239627 |
| ACC 6h | 66.1603 | 33.0007 | 1.46998 | 1.07057 |
| BABA 6h | 47.9301 | 14.3937 | 1.25783 | 0.507307 |
| Chitin 6h | 17.6144 | 3.54147 | 0.540939 | 0.111914 |
| Epi 6h | 19.867 | 3.71212 | 0.581266 | 0.108678 |
| SA 6h | 17.4003 | 3.40695 | 0.570033 | 0.118367 |
| Me-JA 6h | 26.8926 | 10.9091 | 0.654787 | 0.626859 |
Source Transcript PGSC0003DMT400038285 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G39030.1 | +1 | 5e-35 | 128 | 83/221 (38%) | Acyl-CoA N-acyltransferases (NAT) superfamily protein | chr2:16298260-16298946 FORWARD LENGTH=228 |
