Probe CUST_46776_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_46776_PI426222305 | JHI_St_60k_v1 | DMT400038284 | GGGAGCTGTTAGTAGAGAATGGCTTTTTGTGTGCTTAATTGTGCAATTTATTAGTATCTA |
All Microarray Probes Designed to Gene DMG400014772
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_46776_PI426222305 | JHI_St_60k_v1 | DMT400038284 | GGGAGCTGTTAGTAGAGAATGGCTTTTTGTGTGCTTAATTGTGCAATTTATTAGTATCTA |
Microarray Signals from CUST_46776_PI426222305
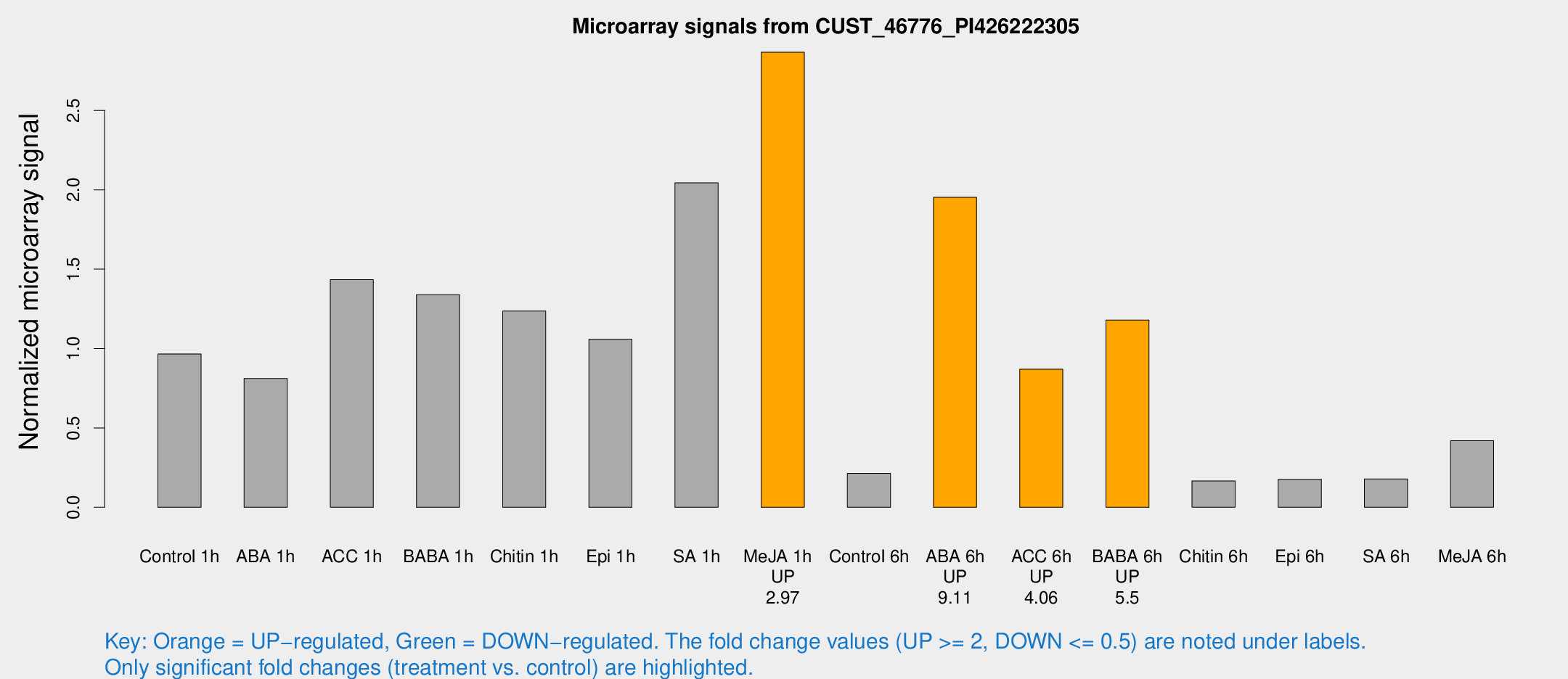
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 4811.61 | 278.625 | 0.965685 | 0.0557587 |
| ABA 1h | 3613.21 | 409.432 | 0.811061 | 0.0468338 |
| ACC 1h | 7828.61 | 1959.46 | 1.43447 | 0.297561 |
| BABA 1h | 6691.43 | 1256.38 | 1.33947 | 0.145044 |
| Chitin 1h | 5530.35 | 319.811 | 1.23585 | 0.144447 |
| Epi 1h | 4557.47 | 263.519 | 1.05829 | 0.0611061 |
| SA 1h | 11060.7 | 2798.42 | 2.04355 | 0.495057 |
| Me-JA 1h | 11772 | 1325.79 | 2.86647 | 0.165498 |
| Control 6h | 1261.58 | 489.862 | 0.214193 | 0.0772731 |
| ABA 6h | 11183.8 | 3125.19 | 1.95224 | 0.471451 |
| ACC 6h | 5404.57 | 1627.16 | 0.869836 | 0.347077 |
| BABA 6h | 7440.46 | 2484.16 | 1.1789 | 0.469731 |
| Chitin 6h | 879.291 | 51.4918 | 0.165658 | 0.0121916 |
| Epi 6h | 1009.98 | 143.46 | 0.176265 | 0.0144927 |
| SA 6h | 899.9 | 139.835 | 0.17837 | 0.0116602 |
| Me-JA 6h | 2150.65 | 373.119 | 0.419706 | 0.0921801 |
Source Transcript PGSC0003DMT400038284 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G39030.1 | +2 | 3e-42 | 148 | 86/238 (36%) | Acyl-CoA N-acyltransferases (NAT) superfamily protein | chr2:16298260-16298946 FORWARD LENGTH=228 |
