Probe CUST_46762_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_46762_PI426222305 | JHI_St_60k_v1 | DMT400038286 | GGTGAAAATCTTCAAAAGTATGCTGATAACAAGGGGGAAATTAAGGAAGCGAACTGTTAG |
All Microarray Probes Designed to Gene DMG400014774
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_46762_PI426222305 | JHI_St_60k_v1 | DMT400038286 | GGTGAAAATCTTCAAAAGTATGCTGATAACAAGGGGGAAATTAAGGAAGCGAACTGTTAG |
Microarray Signals from CUST_46762_PI426222305
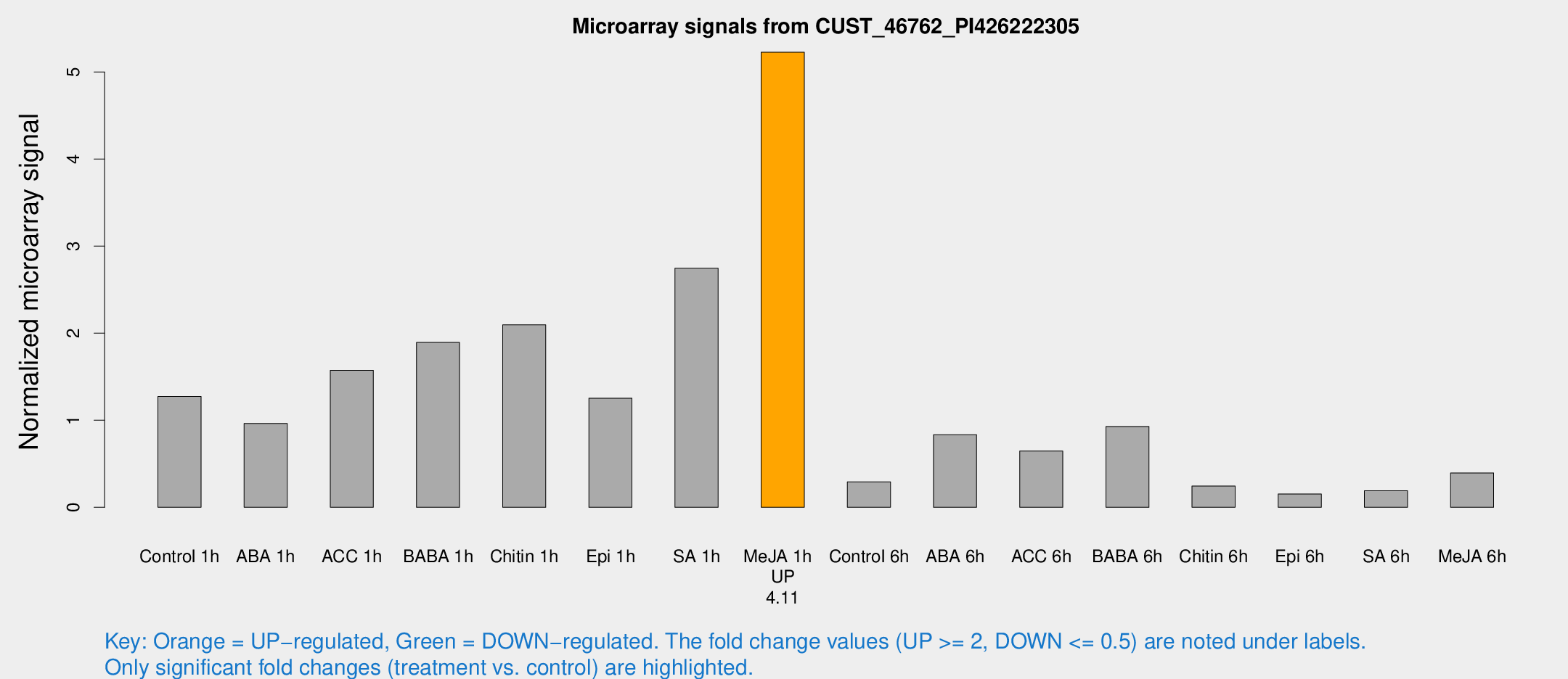
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 428.805 | 82.0648 | 1.27265 | 0.164425 |
| ABA 1h | 294.781 | 76.4109 | 0.962121 | 0.306628 |
| ACC 1h | 599.266 | 179.511 | 1.57333 | 0.529521 |
| BABA 1h | 605.858 | 78.3979 | 1.89383 | 0.109882 |
| Chitin 1h | 616.363 | 36.4871 | 2.09626 | 0.282277 |
| Epi 1h | 357.308 | 37.4266 | 1.25349 | 0.104909 |
| SA 1h | 981.026 | 262.164 | 2.74613 | 0.705435 |
| Me-JA 1h | 1401.54 | 128.016 | 5.22732 | 0.30204 |
| Control 6h | 131.672 | 75.5338 | 0.293071 | 0.169289 |
| ABA 6h | 310.752 | 82.5814 | 0.834418 | 0.186063 |
| ACC 6h | 247.283 | 39.6507 | 0.64594 | 0.169627 |
| BABA 6h | 367.965 | 105.87 | 0.928234 | 0.272014 |
| Chitin 6h | 85.0058 | 6.08071 | 0.244118 | 0.0175146 |
| Epi 6h | 58.8807 | 12.924 | 0.1533 | 0.0312175 |
| SA 6h | 65.7484 | 17.0014 | 0.189206 | 0.0337717 |
| Me-JA 6h | 130.704 | 19.4634 | 0.394407 | 0.0274071 |
Source Transcript PGSC0003DMT400038286 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G39030.1 | +1 | 3e-28 | 110 | 81/291 (28%) | Acyl-CoA N-acyltransferases (NAT) superfamily protein | chr2:16298260-16298946 FORWARD LENGTH=228 |
