Probe CUST_46696_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_46696_PI426222305 | JHI_St_60k_v1 | DMT400024348 | GGTATGTACATAAAATGTGTGAATACCTTCAAGGCAAAGGAAGTTGTAAGTGTAGAGTAA |
All Microarray Probes Designed to Gene DMG401009415
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_46693_PI426222305 | JHI_St_60k_v1 | DMT400024347 | TCTTGGCTGAGCTTACTATTATCAATCGTCTTCGACATAAACATCTTGTCAAATTGCTTG |
| CUST_46696_PI426222305 | JHI_St_60k_v1 | DMT400024348 | GGTATGTACATAAAATGTGTGAATACCTTCAAGGCAAAGGAAGTTGTAAGTGTAGAGTAA |
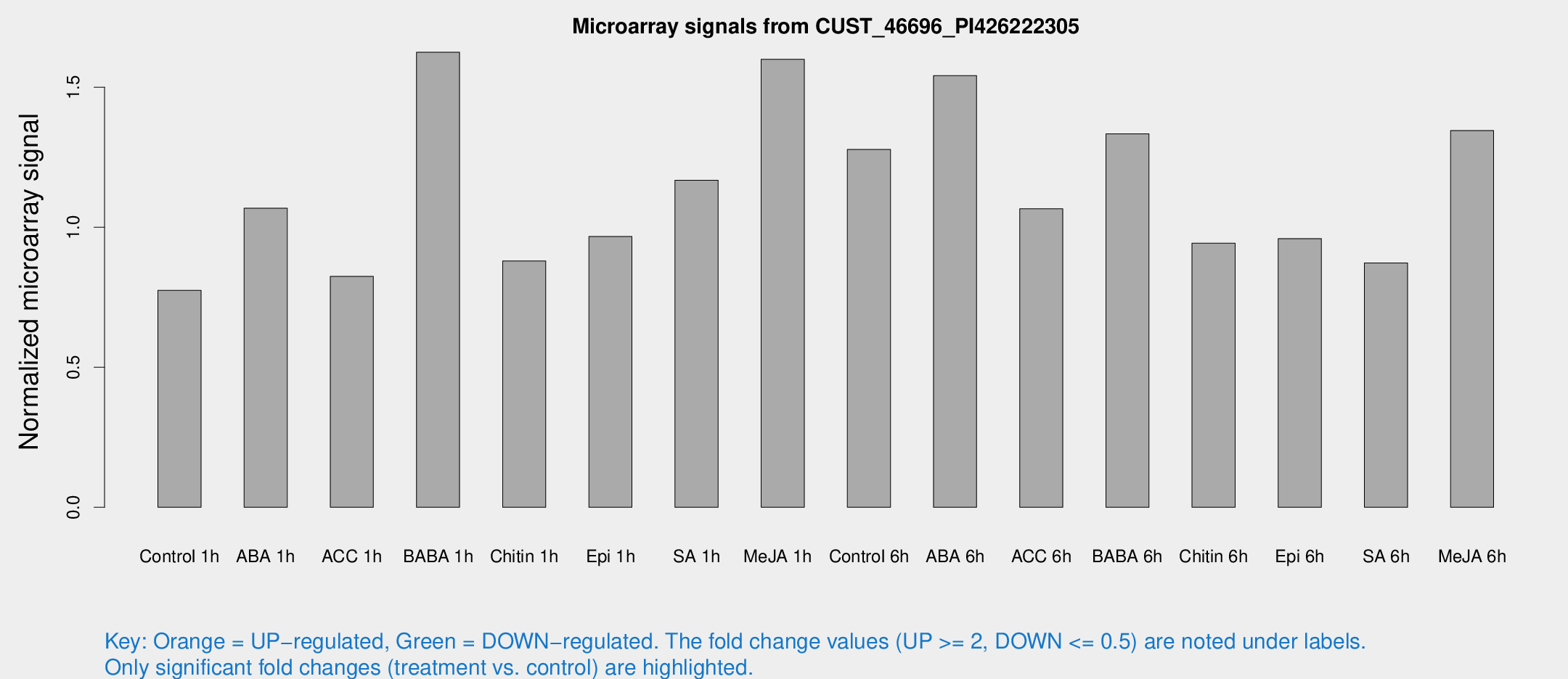
Microarray Signals from CUST_46696_PI426222305
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 5.47093 | 3.17025 | 0.774717 | 0.448718 |
| ABA 1h | 6.75858 | 3.12434 | 1.06778 | 0.511555 |
| ACC 1h | 5.98503 | 3.48028 | 0.824297 | 0.478548 |
| BABA 1h | 11.2175 | 3.45509 | 1.62483 | 0.520964 |
| Chitin 1h | 5.593 | 3.25093 | 0.87978 | 0.508881 |
| Epi 1h | 5.93139 | 3.23115 | 0.966709 | 0.5292 |
| SA 1h | 9.03951 | 3.24733 | 1.16741 | 0.481784 |
| Me-JA 1h | 10.0726 | 3.35088 | 1.59987 | 0.640315 |
| Control 6h | 11.0858 | 5.23136 | 1.27754 | 0.601345 |
| ABA 6h | 13.214 | 4.75386 | 1.54134 | 0.566401 |
| ACC 6h | 9.00814 | 4.14228 | 1.06582 | 0.518934 |
| BABA 6h | 11.3517 | 3.84079 | 1.33314 | 0.525313 |
| Chitin 6h | 7.17444 | 3.69102 | 0.942788 | 0.49706 |
| Epi 6h | 7.71038 | 3.89071 | 0.959482 | 0.493144 |
| SA 6h | 6.09715 | 3.53806 | 0.87267 | 0.506109 |
| Me-JA 6h | 9.76003 | 3.4253 | 1.34529 | 0.5127 |
Source Transcript PGSC0003DMT400024348 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G06740.1 | +3 | 3e-53 | 194 | 127/364 (35%) | Concanavalin A-like lectin protein kinase family protein | chr5:2084094-2086052 FORWARD LENGTH=652 |
