Probe CUST_46438_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_46438_PI426222305 | JHI_St_60k_v1 | DMT400005893 | GAAGATATTCCTTCACCTACTAGCAGGGAAGAAAATACTTATGGTTTTTGAATCCTTATG |
All Microarray Probes Designed to Gene DMG400002291
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_46438_PI426222305 | JHI_St_60k_v1 | DMT400005893 | GAAGATATTCCTTCACCTACTAGCAGGGAAGAAAATACTTATGGTTTTTGAATCCTTATG |
Microarray Signals from CUST_46438_PI426222305
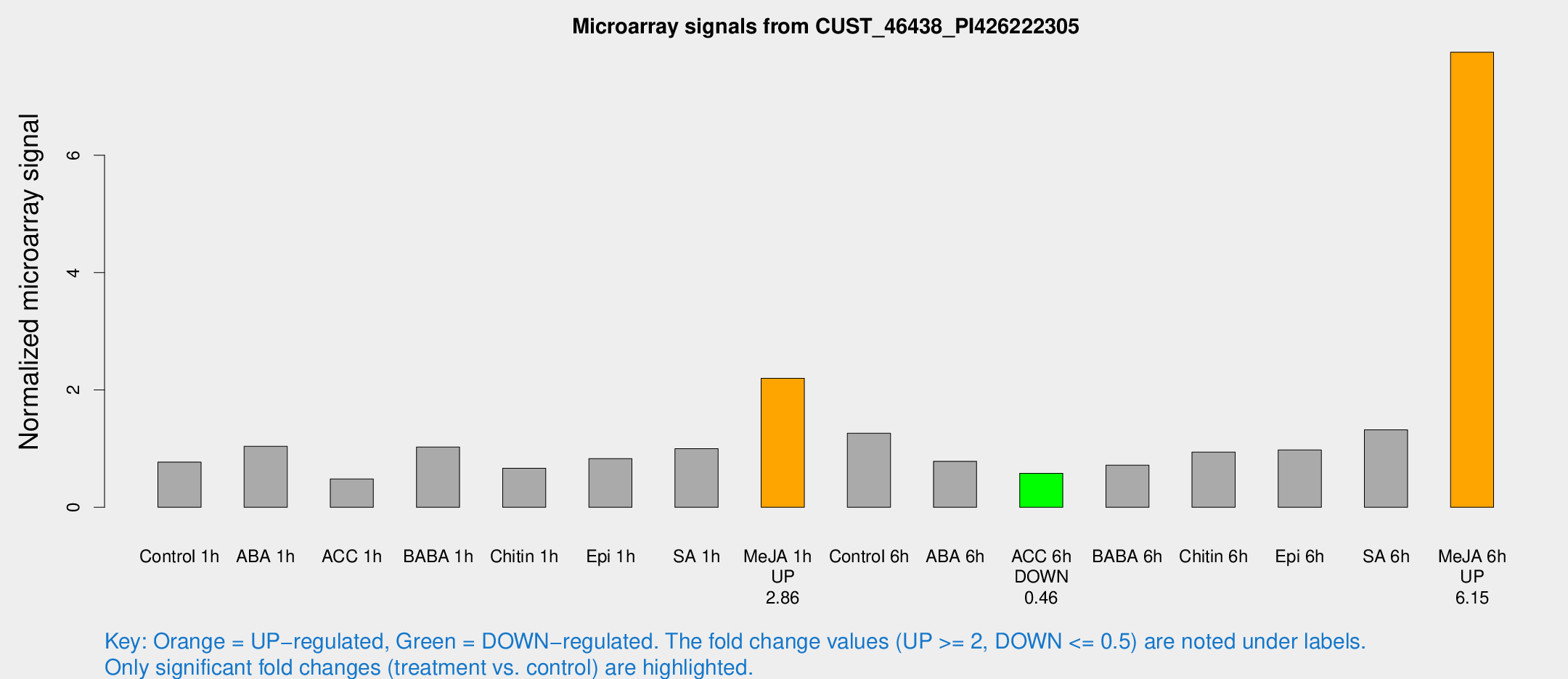
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 309.922 | 51.4337 | 0.77033 | 0.161576 |
| ABA 1h | 371.019 | 68.998 | 1.03945 | 0.120873 |
| ACC 1h | 225.802 | 78.9987 | 0.481962 | 0.185656 |
| BABA 1h | 395.104 | 58.6454 | 1.02653 | 0.0640835 |
| Chitin 1h | 234.492 | 14.1244 | 0.666416 | 0.0540188 |
| Epi 1h | 284.325 | 35.1257 | 0.829996 | 0.1032 |
| SA 1h | 432.848 | 123.862 | 1.00063 | 0.273762 |
| Me-JA 1h | 712.819 | 96.5953 | 2.20019 | 0.127383 |
| Control 6h | 497.361 | 48.1374 | 1.26112 | 0.151142 |
| ABA 6h | 330.535 | 44.4623 | 0.784051 | 0.117235 |
| ACC 6h | 266.586 | 41.6304 | 0.580232 | 0.122383 |
| BABA 6h | 332.998 | 80.6419 | 0.71729 | 0.203917 |
| Chitin 6h | 391.179 | 22.9044 | 0.939387 | 0.0821298 |
| Epi 6h | 436.603 | 53.8006 | 0.977229 | 0.186407 |
| SA 6h | 523.548 | 85.4927 | 1.32052 | 0.109374 |
| Me-JA 6h | 3040.58 | 289.257 | 7.75694 | 1.33198 |
Source Transcript PGSC0003DMT400005893 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G30370.1 | +2 | 2e-177 | 514 | 270/488 (55%) | alpha/beta-Hydrolases superfamily protein | chr1:10719169-10720758 REVERSE LENGTH=529 |
