Probe CUST_46305_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_46305_PI426222305 | JHI_St_60k_v1 | DMT400034124 | CCAGAGAAGATAAAAGACACAACAATTGTGTATATGTCACAACTCACAACTTGAAGACTT |
All Microarray Probes Designed to Gene DMG401013112
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_46305_PI426222305 | JHI_St_60k_v1 | DMT400034124 | CCAGAGAAGATAAAAGACACAACAATTGTGTATATGTCACAACTCACAACTTGAAGACTT |
| CUST_46296_PI426222305 | JHI_St_60k_v1 | DMT400034138 | CCTTTGGACAGGTAAACAGACAGTTCAATTTAGTCACAATTTATACATACTCTGGAGTTA |
Microarray Signals from CUST_46305_PI426222305
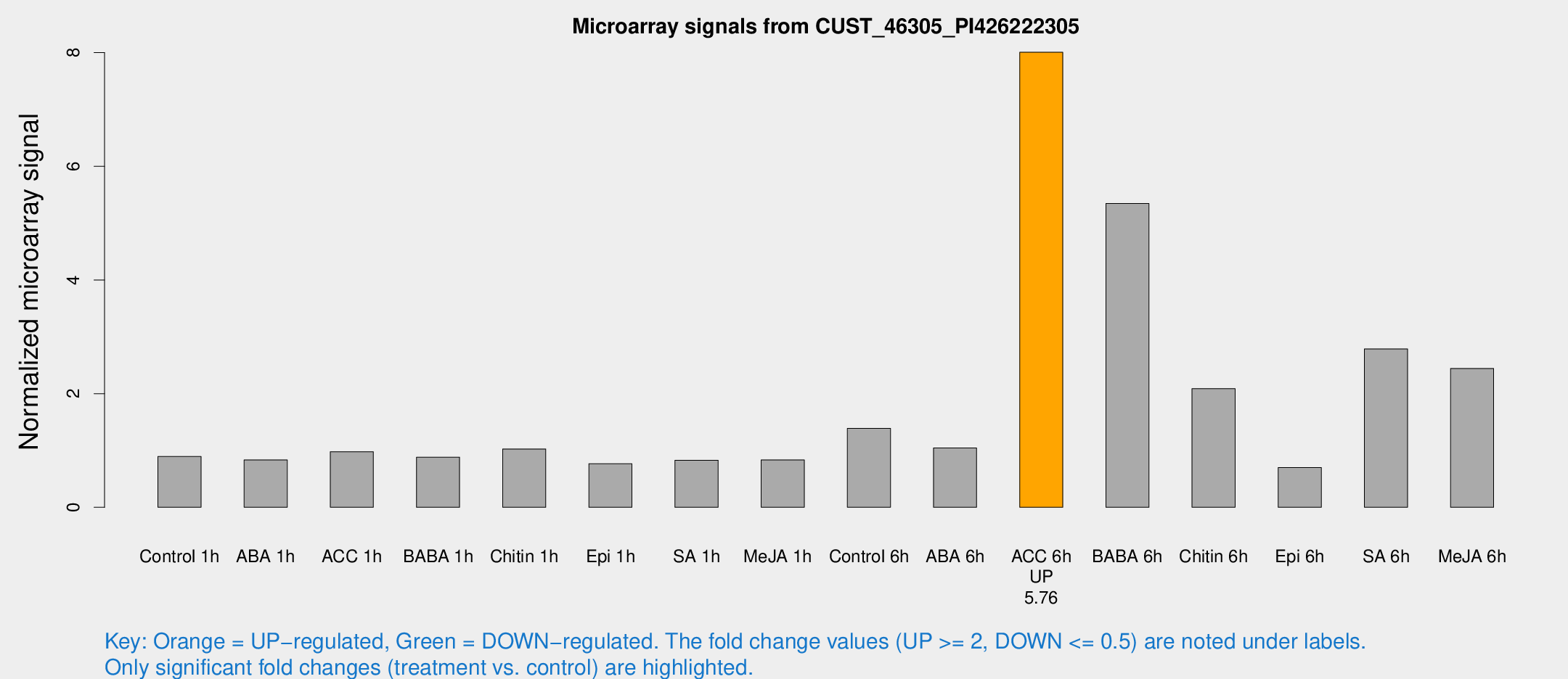
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 7.65285 | 3.13347 | 0.894262 | 0.418784 |
| ABA 1h | 6.10047 | 3.02759 | 0.832446 | 0.431277 |
| ACC 1h | 8.6405 | 3.44399 | 0.979036 | 0.438777 |
| BABA 1h | 6.95858 | 3.29172 | 0.881731 | 0.430487 |
| Chitin 1h | 7.65599 | 3.18743 | 1.02668 | 0.451404 |
| Epi 1h | 5.29973 | 3.07657 | 0.766562 | 0.438234 |
| SA 1h | 7.12809 | 3.1106 | 0.828989 | 0.384402 |
| Me-JA 1h | 5.51398 | 3.21204 | 0.832539 | 0.482648 |
| Control 6h | 12.3768 | 3.4074 | 1.38998 | 0.494994 |
| ABA 6h | 10.5564 | 4.48755 | 1.04354 | 0.465441 |
| ACC 6h | 75.1937 | 8.38082 | 8.00736 | 1.13276 |
| BABA 6h | 57.4791 | 24.2522 | 5.3463 | 2.36656 |
| Chitin 6h | 21.5237 | 9.55244 | 2.08915 | 1.06985 |
| Epi 6h | 6.43893 | 3.77787 | 0.699844 | 0.405999 |
| SA 6h | 26.8988 | 9.86398 | 2.78524 | 1.0964 |
| Me-JA 6h | 19.9752 | 3.50549 | 2.44347 | 0.444015 |
Source Transcript PGSC0003DMT400034124 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G15520.1 | +1 | 0.0 | 2026 | 1003/1408 (71%) | pleiotropic drug resistance 12 | chr1:5331993-5338175 REVERSE LENGTH=1423 |
