Probe CUST_46190_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_46190_PI426222305 | JHI_St_60k_v1 | DMT400011423 | AGTGTTTTATTTTGAGAAAATGGATAGTAGTTTACTGATTGCTGCTCCCTCTATGTGACA |
All Microarray Probes Designed to Gene DMG400004479
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_46190_PI426222305 | JHI_St_60k_v1 | DMT400011423 | AGTGTTTTATTTTGAGAAAATGGATAGTAGTTTACTGATTGCTGCTCCCTCTATGTGACA |
Microarray Signals from CUST_46190_PI426222305
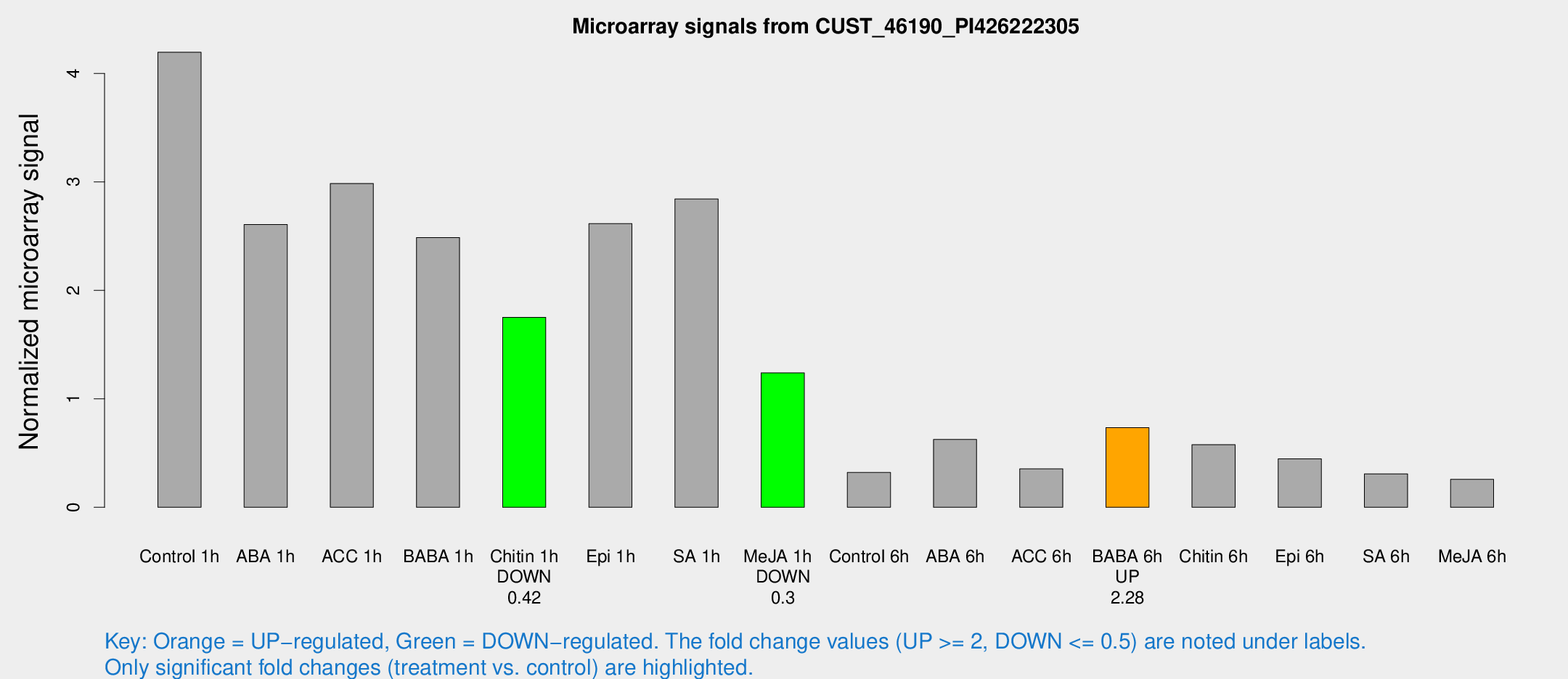
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 1656.41 | 95.9335 | 4.19545 | 0.242405 |
| ABA 1h | 929.592 | 132.804 | 2.60742 | 0.285721 |
| ACC 1h | 1393.21 | 438.839 | 2.98486 | 1.08768 |
| BABA 1h | 1002.76 | 221.772 | 2.48727 | 0.384074 |
| Chitin 1h | 621.759 | 36.1715 | 1.75147 | 0.122251 |
| Epi 1h | 892.994 | 51.7736 | 2.61495 | 0.151346 |
| SA 1h | 1159.56 | 104.147 | 2.84212 | 0.164345 |
| Me-JA 1h | 402.399 | 39.974 | 1.23945 | 0.0725254 |
| Control 6h | 128.208 | 13.0944 | 0.321407 | 0.0209067 |
| ABA 6h | 268.017 | 38.3462 | 0.625791 | 0.0655329 |
| ACC 6h | 160.151 | 10.3666 | 0.35429 | 0.0508989 |
| BABA 6h | 326.306 | 28.3323 | 0.734358 | 0.0433998 |
| Chitin 6h | 249.564 | 41.5417 | 0.577259 | 0.102043 |
| Epi 6h | 198.313 | 12.2992 | 0.446557 | 0.0314029 |
| SA 6h | 125.784 | 24.4369 | 0.308629 | 0.0320956 |
| Me-JA 6h | 112.543 | 38.2435 | 0.257705 | 0.0656287 |
Source Transcript PGSC0003DMT400011423 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G32870.1 | +1 | 4e-39 | 135 | 67/151 (44%) | Polyketide cyclase/dehydrase and lipid transport superfamily protein | chr4:15862168-15862641 FORWARD LENGTH=157 |
