Probe CUST_46151_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_46151_PI426222305 | JHI_St_60k_v1 | DMT400018498 | GAATGGGCCCAAATTTATGGGCCAATTTTTTCTTTTTACCTTGGGTCACAACTAAACGTG |
All Microarray Probes Designed to Gene DMG400007181
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_46151_PI426222305 | JHI_St_60k_v1 | DMT400018498 | GAATGGGCCCAAATTTATGGGCCAATTTTTTCTTTTTACCTTGGGTCACAACTAAACGTG |
Microarray Signals from CUST_46151_PI426222305
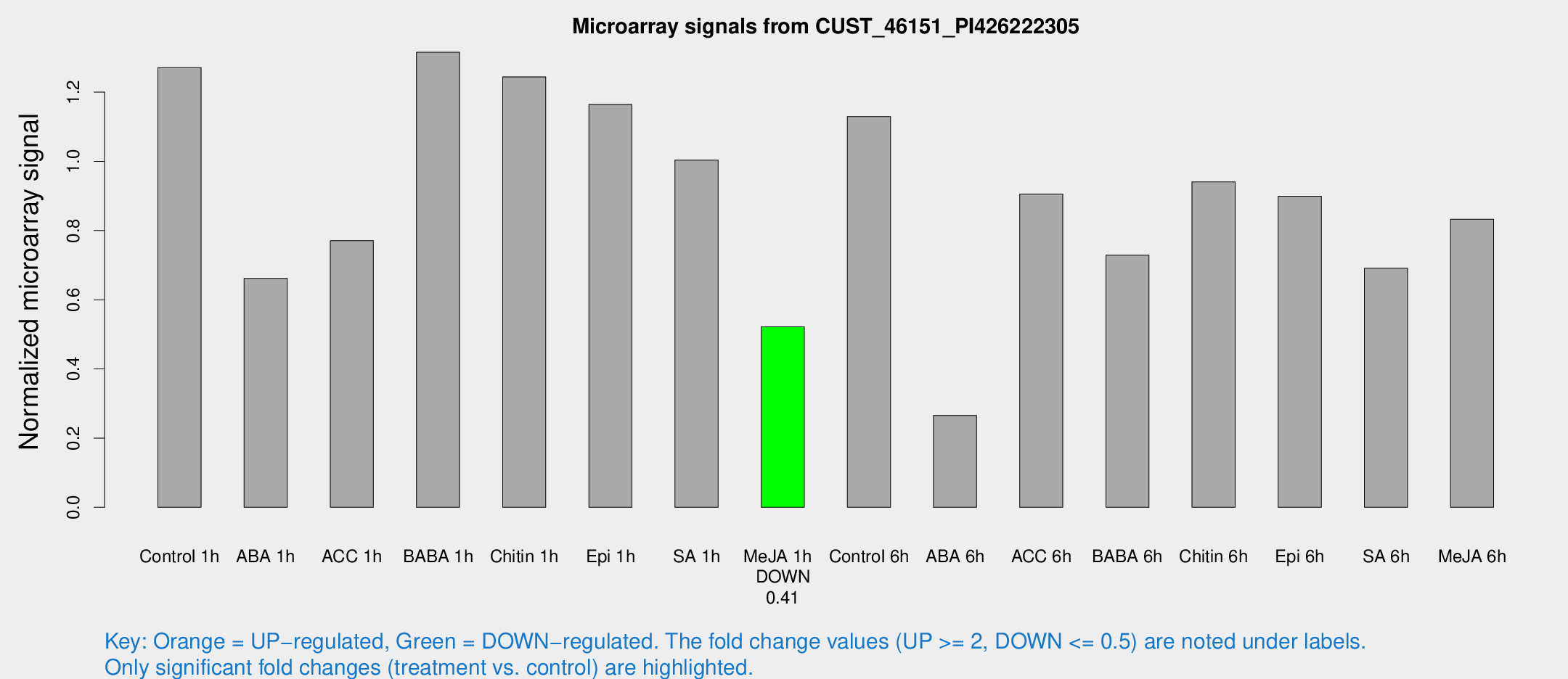
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 325.63 | 65.0509 | 1.27109 | 0.163417 |
| ABA 1h | 146.925 | 18.9982 | 0.661834 | 0.0421028 |
| ACC 1h | 215.16 | 57.6353 | 0.770964 | 0.204952 |
| BABA 1h | 328.758 | 72.3105 | 1.31556 | 0.187684 |
| Chitin 1h | 276.864 | 16.3402 | 1.2443 | 0.0732087 |
| Epi 1h | 250.071 | 18.5999 | 1.16462 | 0.068817 |
| SA 1h | 259.158 | 36.7777 | 1.00366 | 0.0935969 |
| Me-JA 1h | 105.965 | 9.80819 | 0.521758 | 0.0419742 |
| Control 6h | 300.681 | 73.1064 | 1.12949 | 0.22025 |
| ABA 6h | 75.9398 | 21.2808 | 0.265622 | 0.0683778 |
| ACC 6h | 262.486 | 38.9275 | 0.905735 | 0.0805564 |
| BABA 6h | 215.671 | 50.3161 | 0.728795 | 0.183521 |
| Chitin 6h | 256.621 | 46.043 | 0.940704 | 0.198127 |
| Epi 6h | 279.669 | 82.5041 | 0.899199 | 0.418385 |
| SA 6h | 190.721 | 55.9595 | 0.691204 | 0.368032 |
| Me-JA 6h | 234.698 | 76.3442 | 0.832554 | 0.278737 |
Source Transcript PGSC0003DMT400018498 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G40890.1 | +2 | 3e-120 | 360 | 169/290 (58%) | cytochrome P450, family 98, subfamily A, polypeptide 3 | chr2:17058291-17060532 REVERSE LENGTH=508 |
