Probe CUST_45890_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_45890_PI426222305 | JHI_St_60k_v1 | DMT400048001 | GACACTGGTGTTAAAATTAGCTTCTTCAATCCTCAGTAACTAGTATGAACATGCGAAATT |
All Microarray Probes Designed to Gene DMG400018651
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_45938_PI426222305 | JHI_St_60k_v1 | DMT400048002 | TTTTGCTTTCAGAGGCCACAATCTCATACTTGAGATTCAGGCGACAATGCCTTCGAGTGA |
| CUST_45890_PI426222305 | JHI_St_60k_v1 | DMT400048001 | GACACTGGTGTTAAAATTAGCTTCTTCAATCCTCAGTAACTAGTATGAACATGCGAAATT |
Microarray Signals from CUST_45890_PI426222305
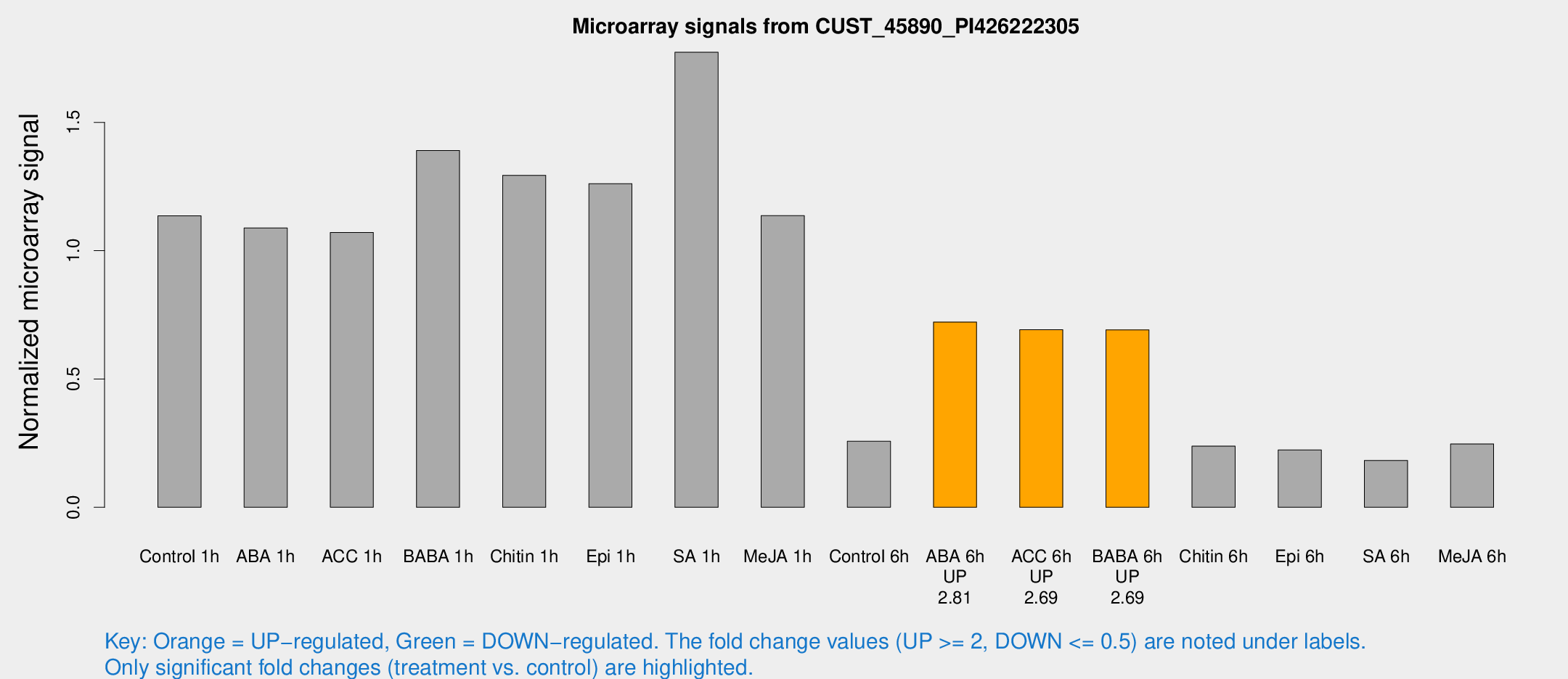
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 13260.8 | 767.424 | 1.13623 | 0.065601 |
| ABA 1h | 11336 | 1178.59 | 1.08887 | 0.0628667 |
| ACC 1h | 15073.9 | 4916.52 | 1.07068 | 0.433924 |
| BABA 1h | 16785.7 | 4013.81 | 1.39046 | 0.255713 |
| Chitin 1h | 13602.4 | 880.387 | 1.29387 | 0.0923841 |
| Epi 1h | 12719.8 | 734.939 | 1.26131 | 0.0728225 |
| SA 1h | 22070.6 | 4205.25 | 1.77345 | 0.256229 |
| Me-JA 1h | 11002.7 | 1541.36 | 1.13648 | 0.0706816 |
| Control 6h | 3240.53 | 788.297 | 0.257149 | 0.0526392 |
| ABA 6h | 9814.45 | 2701.68 | 0.722069 | 0.20568 |
| ACC 6h | 9347.84 | 968.567 | 0.692474 | 0.0548714 |
| BABA 6h | 9864.02 | 3118.07 | 0.691658 | 0.189397 |
| Chitin 6h | 3048.64 | 507.328 | 0.238415 | 0.0465352 |
| Epi 6h | 3018.43 | 467.903 | 0.223673 | 0.0242427 |
| SA 6h | 2443.15 | 828.736 | 0.182435 | 0.0585498 |
| Me-JA 6h | 3132.4 | 902.509 | 0.246819 | 0.0568123 |
Source Transcript PGSC0003DMT400048001 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G47670.1 | +3 | 3e-81 | 256 | 110/207 (53%) | Transmembrane amino acid transporter family protein | chr1:17536834-17539486 REVERSE LENGTH=519 |
