Probe CUST_45278_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_45278_PI426222305 | JHI_St_60k_v1 | DMT400027594 | CACTAGTGATACTGGACCTTATTGGGAATCACCAAGAAAATTGGTTGATGAAAAGTATAA |
All Microarray Probes Designed to Gene DMG400010638
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_45278_PI426222305 | JHI_St_60k_v1 | DMT400027594 | CACTAGTGATACTGGACCTTATTGGGAATCACCAAGAAAATTGGTTGATGAAAAGTATAA |
Microarray Signals from CUST_45278_PI426222305
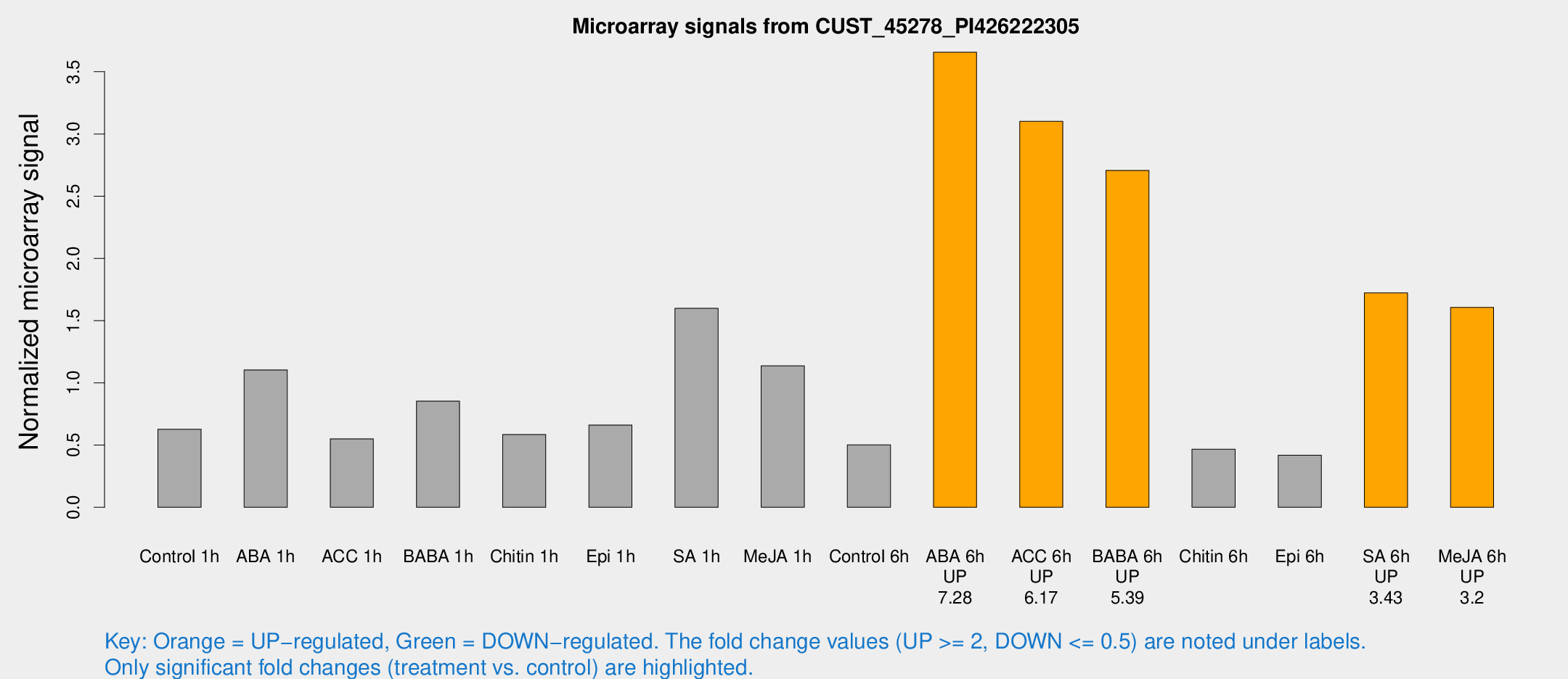
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 565.24 | 68.1343 | 0.627674 | 0.0940158 |
| ABA 1h | 884.928 | 123.759 | 1.10418 | 0.112928 |
| ACC 1h | 624.612 | 289.065 | 0.550445 | 0.262475 |
| BABA 1h | 793.399 | 215.154 | 0.852964 | 0.18012 |
| Chitin 1h | 470.078 | 46.1339 | 0.584287 | 0.0340391 |
| Epi 1h | 517.023 | 74.4032 | 0.66104 | 0.0965309 |
| SA 1h | 1564.66 | 403.653 | 1.59895 | 0.40496 |
| Me-JA 1h | 857.292 | 165.878 | 1.13734 | 0.140692 |
| Control 6h | 475.988 | 108.919 | 0.50224 | 0.0877755 |
| ABA 6h | 3952.41 | 1215.82 | 3.65651 | 1.43381 |
| ACC 6h | 3226.59 | 509.585 | 3.10082 | 0.809628 |
| BABA 6h | 2884.76 | 743.257 | 2.70653 | 0.762306 |
| Chitin 6h | 445.063 | 43.0999 | 0.467041 | 0.0626092 |
| Epi 6h | 430.381 | 69.5978 | 0.418506 | 0.0411685 |
| SA 6h | 1608.82 | 366.615 | 1.72262 | 0.258758 |
| Me-JA 6h | 1436.47 | 169.215 | 1.60616 | 0.310427 |
Source Transcript PGSC0003DMT400027594 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G41380.1 | +3 | 2e-113 | 333 | 162/264 (61%) | S-adenosyl-L-methionine-dependent methyltransferases superfamily protein | chr2:17251981-17252886 FORWARD LENGTH=269 |
