Probe CUST_45225_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_45225_PI426222305 | JHI_St_60k_v1 | DMT400010863 | CTTGTTGTCTCTCAAACCATCAATAACTCTCAGAACGTTGGAGCTTAAAAGAAATATAGT |
All Microarray Probes Designed to Gene DMG400004252
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_45225_PI426222305 | JHI_St_60k_v1 | DMT400010863 | CTTGTTGTCTCTCAAACCATCAATAACTCTCAGAACGTTGGAGCTTAAAAGAAATATAGT |
| CUST_45198_PI426222305 | JHI_St_60k_v1 | DMT400010864 | GGCTCAGATTGTCAACTGGACCTTTTACTCACAATTATGGTCTCAAATTTAGTTAATTTG |
Microarray Signals from CUST_45225_PI426222305
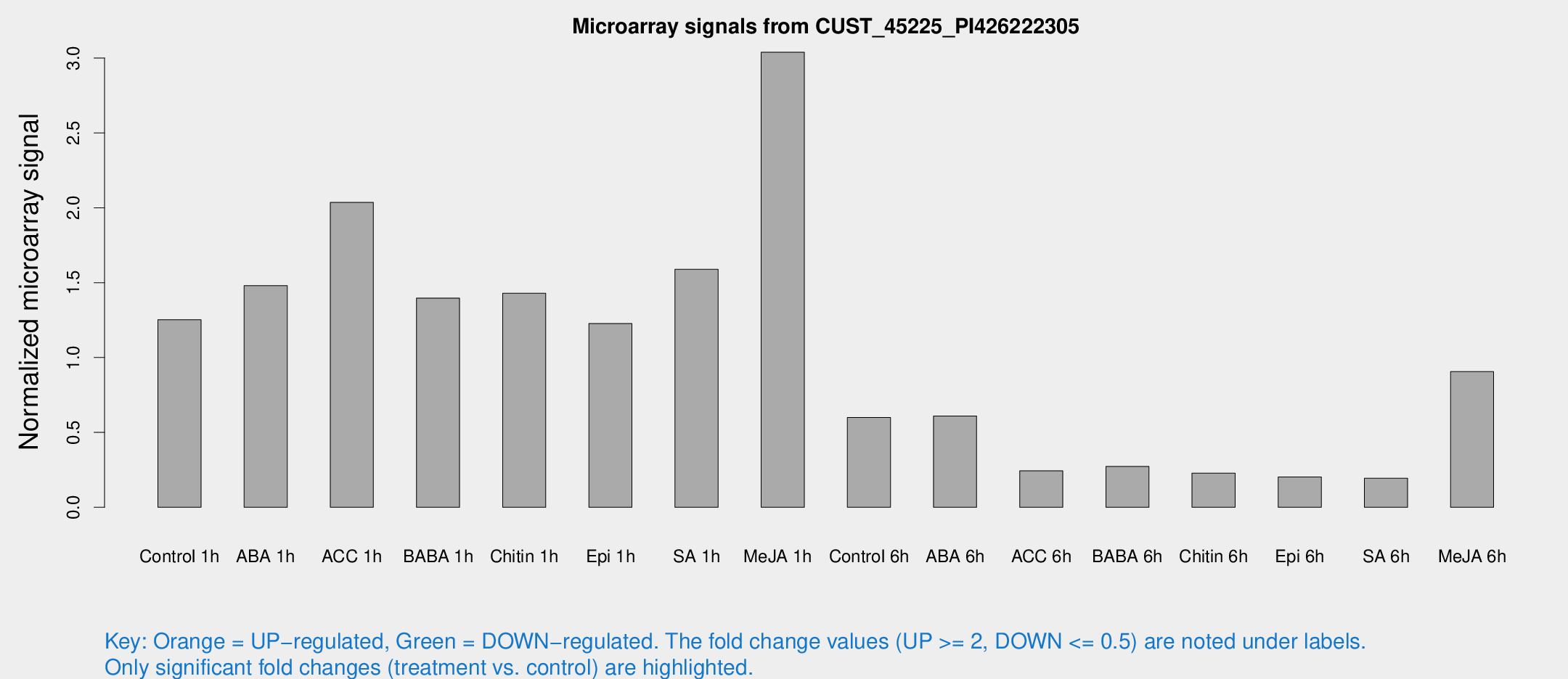
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 658.103 | 152.461 | 1.25258 | 0.206127 |
| ABA 1h | 664.717 | 83.2397 | 1.47966 | 0.113904 |
| ACC 1h | 1061.3 | 131.59 | 2.03572 | 0.247579 |
| BABA 1h | 720.275 | 166.777 | 1.39713 | 0.238086 |
| Chitin 1h | 658.406 | 95.3526 | 1.42891 | 0.137681 |
| Epi 1h | 539.516 | 64.8172 | 1.22682 | 0.129244 |
| SA 1h | 851.456 | 173.778 | 1.5889 | 0.297355 |
| Me-JA 1h | 1285.88 | 224.675 | 3.03846 | 0.364023 |
| Control 6h | 358.4 | 156.475 | 0.599864 | 0.224079 |
| ABA 6h | 342.948 | 77.9331 | 0.609486 | 0.117151 |
| ACC 6h | 140.567 | 11.0805 | 0.243301 | 0.0378214 |
| BABA 6h | 157.481 | 26.0831 | 0.272481 | 0.0564222 |
| Chitin 6h | 124.531 | 20.6546 | 0.227727 | 0.0404959 |
| Epi 6h | 124.714 | 34.7905 | 0.202226 | 0.0792235 |
| SA 6h | 115.377 | 42.0618 | 0.193587 | 0.0752207 |
| Me-JA 6h | 454.436 | 38.1977 | 0.905843 | 0.0595925 |
Source Transcript PGSC0003DMT400010863 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G22920.1 | +3 | 5e-109 | 322 | 165/264 (63%) | non-yellowing 1 | chr4:12016776-12017969 REVERSE LENGTH=268 |
